Single-Cell Spatial Transcriptomics: Spatial Visualization Guide
Module Introduction
This tutorial is specifically designed to handle the underlying spatial geometric transformations for SeekGene single-cell spatial transcriptomics products. It aims to help you deeply understand and achieve precise registration between spatial transcriptomic chip coordinates and histological tissue section images.
Before diving into the code, it is essential to understand the spatial coordinate system of single-cell spatial transcriptomics products. In the output matrices from the analysis pipeline, spatial coordinates (often referred to as "chip pixels" or "spot coordinates") are established based on the sequencing chip's internal grid. This fundamentally differs from the optical image pixels we typically handle; it intrinsically represents the underlying physical resolution of the chip. For instance, in SeekGene high-resolution chips, 1 chip pixel roughly corresponds to a true physical distance of 0.2653 micrometers (μm).
In real-world bioinformatics analysis, researchers inevitably need to perform geometric transformations on these raw coordinates—whether to seamlessly integrate gene expression profiles with tissue morphology or to accurately measure physical distances within the cellular microenvironment. If your focus is on the spatial visualization of downstream analysis results, such as biological clustering or cell type annotation, please refer to other tutorials in the single-cell spatial transcriptomics series. This tutorial, however, focuses on the fundamental groundwork, specifically breaking down and demonstrating the two core spatial transformation tasks involved in coordinate mapping and image registration:
- 1. Conversion and Unification of Spatial Coordinates: As the foundation of spatial visualization, this primarily involves transformations across two dimensions:
- Coordinate to Physical Distance: Converting abstract relative coordinates into true physical distances (μm) with actual biological significance, based on the chip's physical resolution.
- Coordinate to Pixel Mapping: Establishing a mathematical mapping between the physical coordinate system of the transcriptomic data and the optical pixel coordinate system of the histological image.
- 2. Interactive High-Precision Image Alignment (Interactive Affine Registration): Following the initial coordinate mapping, this tutorial provides an offline HTML interactive alignment tool to correct for translation or rotation artifacts introduced during tissue sectioning and mounting. You can perform visual fine-tuning—such as translation, scaling, and rotation—via a web interface, thereby achieving lossless rigid registration of the full-resolution original image.
Input File Preparation
This module requires the following input files:
- Required Cell Spatial Localization File: Must be the
_filtered_feature_bc_matrix/cell_locations.tsv.gzfile generated by the standard bioinformatics pipeline. This file contains the physical grid coordinates of each segmented cell on the sequencing chip. - Required Histological Tissue Section Image: Must be an image file that has undergone quality control (QC) and initial alignment via SeekSpaceTools, typically named
*_aligned_HE.pngor*_aligned_DAPI.png. This histological image serves as the basal reference for spatial visualization, ensuring a preliminary spatial concordance with the transcriptomic data.
Import Dependencies
import pandas as pd
import numpy as np
from PIL import Image
import matplotlib.pyplot as plt
import os, base64, json
import cv2
import io
import plotly# === Global Parameter Configuration ===
# Physical data boundaries of the sequencing chip
# Note:
# In SeekSpaceTools v1.0.2, the chip's physical boundary is fixed at 55050x19906.
# For software versions v1.0.3 and above, the chip's physical boundary is typically 56500x20434.
# Specific boundary parameters can be extracted from the .rds or .h5ad objects.
CHIP_MAX_X = 55050
CHIP_MAX_Y = 19906
# Chip physical resolution (micrometers per unit coordinate, μm)
RESOLUTION_UM = 0.2653Conversion and Unification of Spatial Coordinates
This section guides you through the foundational tasks of spatial visualization: converting chip grid coordinates into true physical micrometer (μm) distances, and unifying the transcriptomic data with the image coordinate system at a macroscopic level.
Load Cell Coordinates and Histological Image Data
Prior to any spatial transformation, we must first load two core datasets:
- Cell Spatial Coordinate Data: Typically
cell_locations.tsv.gz, which records the original relative spatial localization (i.e., chip coordinates) of each segmented cell on the sequencing chip grid. - Tissue Section Image: Load the initially aligned H&E or DAPI stained image and extract the shape attributes of its pixel matrix. The image dimensions (width and height) provide critical scale references for defining plotting axis limits and calculating pixel mapping scaling ratios.
table = pd.read_csv("A5_filtered_feature_bc_matrix/cell_locations.tsv.gz", sep="\t")
table.head()| Cell_Barcode | X | Y | |
|---|---|---|---|
| 0 | AAACCCAATGACACGATACTGTG | 35293 | 6684 |
| 1 | AAACCCAATGCGCAGTGCTGCGC | 41981 | 2808 |
| 2 | AAACCCAATGCGGACATACTGTG | 3052 | 11748 |
| 3 | AAACCCAATGCTTCGACTAGTCA | 24843 | 6148 |
| 4 | AAACCCAATGGACCACGTAGTCA | 24279 | 4715 |
img = Image.open('A5_aligned_HE.png')
img_array = np.array(img)
img_array.shapeInitial Coordinate System Overlay and Visualization (Based on Chip Coordinates)
In this step, we directly overlay and visualize the loaded tissue image with the raw chip coordinates of the cells to perform a preliminary alignment check.
Note: If you are using an H&E stained image, it is often derived from an adjacent section (Adjacent Slice) to the one used for spatial transcriptomics sequencing. Due to the continuous 3D morphological variation of biological tissues and independent minor deformations (artifacts) introduced during the preparation of the two sections, slight local mismatches between the tissue contour in the image and the cell coordinates are normal experimental phenomena and do not indicate a coordinate mapping error.
Code Logic Breakdown:
extent=[0, CHIP_MAX_X, 0, CHIP_MAX_Y]: This is a pivotal parameter inmatplotlib.pyplot.imshow. Since our cell scatter coordinates (X, Y) are based on the chip's physical grid (e.g., 0 ~ 55050) while the loaded image is in an optical pixel coordinate system (e.g., height 1447 × width 4000 pixels), their scales are completely disparate. By configuringextent, we forcefully "stretch" the histological image during rendering to span the entire coordinate bounding box of the chip.- Note: At this stage, the image is proportionally scaled and forcefully fitted onto the chip's relative coordinate space. This represents a "forced mapping" strictly at the visual level, primarily utilized to verify whether the macroscopic orientation of the tissue and the cell distribution contours are broadly concordant. It does not yet involve true physical distance conversions.
plt.figure(figsize=(20, 12))
# Plot the background histological image first
ax = plt.gca()
ax.imshow(img,
cmap='gray',
extent=[0, CHIP_MAX_X, 0, CHIP_MAX_Y], # Key: Define the image coordinate extent
aspect='auto',
alpha=1)
# Plot the transcriptomic cell scatter overlay
ax.scatter(
table['X'],
table['Y'],
s=5,
alpha=0.8,
edgecolors='none',
rasterized=True
)
ax.set_aspect('equal')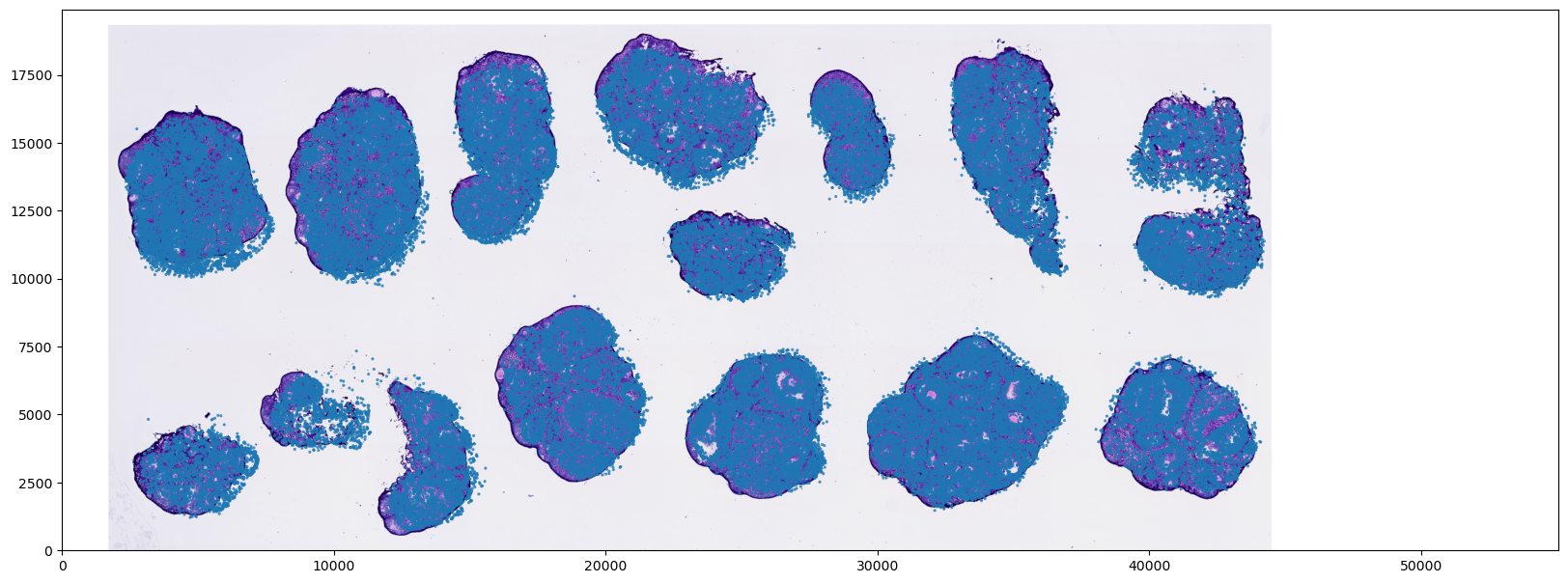
Convert Cell Coordinates to True Physical Distances (μm)
To facilitate downstream spatial analyses with actual biological significance (e.g., calculating true spatial distances within the cellular microenvironment), we must convert the abstract chip coordinates into tangible physical length units (micrometers, μm).
Core Logic:
- Physical Scale Conversion: Multiply the raw cell coordinates by the physical conversion factor (i.e., the true physical distance in μm corresponding to each chip coordinate unit) to derive the physical distance coordinates.
- Visualization and Performance Considerations: When rendering scatter plots based on physical distances, simply adjust the coordinate bounding box of the background image at the plotting API level. It is strongly recommended to avoid direct upsampling of the original image's pixel matrix, as this may trigger memory overflow (OOM) and compromise the high-resolution histological details.
table["X_um"] = table["X"] * RESOLUTION_UM
table["Y_um"] = table["Y"] * RESOLUTION_UMtable["X_um"].describe()mean 6158.961242
std 3163.637123
min 624.781500
25% 3458.981400
50% 6194.755000
75% 8962.629900
max 11734.749600
Name: X_um, dtype: float64
plt.figure(figsize=(20, 12))
# Plot the background histological image first
ax = plt.gca()
ax.imshow(img,
cmap='gray',
extent=[0, CHIP_MAX_X * RESOLUTION_UM, 0, CHIP_MAX_Y * RESOLUTION_UM], # Key: Define the image coordinate extent
aspect='auto',
alpha=1) # Set opacity level
# Plot the transcriptomic cell scatter overlay
ax.scatter(
table['X_um'],
table['Y_um'],
s=5,
alpha=0.8,
edgecolors='none',
rasterized=True
)
ax.set_aspect('equal')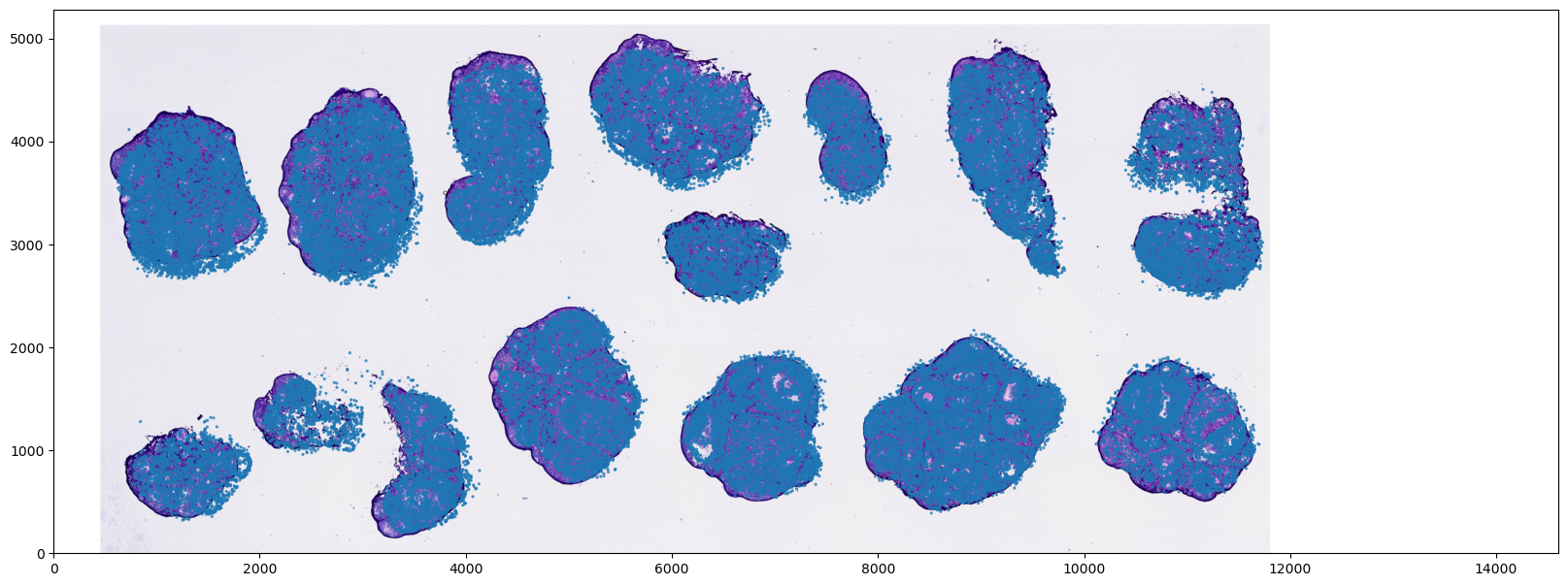
Convert Cell Coordinates to Optical Image Pixels
In specific analytical scenarios (such as extracting localized morphological features from the tissue section), it is necessary to map the spatial coordinates of the transcriptomic cells into the true optical image pixel coordinate system.
Core Logic:
- Proportional Coordinate Scaling: Utilizing the previously calculated scaling ratios, directly downscale the chip coordinates of the cells proportionally to map them into the actual pixel dimensions of the histological image.
- Efficient Registration Strategy: During plotting and alignment, we mathematically transform the "1D coordinate vectors" rather than forcefully resizing the "high-resolution background image." This approach incurs minimal computational overhead, prevents memory issues, and perfectly preserves the morphological features and resolution of the original tissue image.
scale_ratio_x= CHIP_MAX_X/img_array.shape[1]
scale_ratio_y = CHIP_MAX_Y/img_array.shape[0]
print(scale_ratio_x, scale_ratio_y)table["X_img"] = table["X"]/scale_ratio_x
table["Y_img"] = table["Y"]/scale_ratio_yplt.figure(figsize=(20, 12))
# Plot the background histological image first
ax = plt.gca()
ax.imshow(img,
cmap='gray', # Comment this out if providing an H&E stained image
extent=[0, img_array.shape[1], 0, img_array.shape[0]], # Key: Define the image coordinate extent
aspect='auto',
alpha=1) # Set opacity level
# Plot the transcriptomic cell scatter overlay
ax.scatter(
table['X_img'],
table['Y_img'],
s=5,
alpha=0.8,
edgecolors='none',
rasterized=True
)
ax.set_aspect('equal')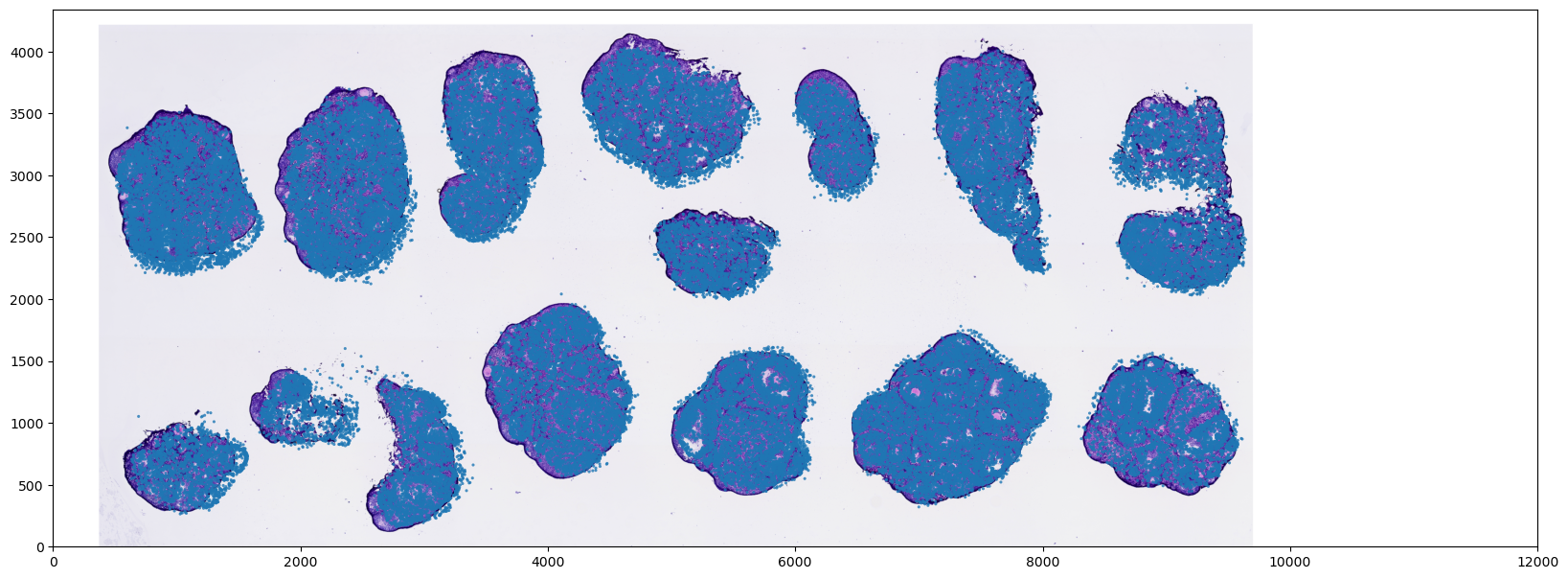
Interactive High-Precision Image Registration
Following the initial coordinate mapping, it is common to observe slight deviations between the tissue image and the transcriptomic cell scatter overlay. To correct physical offsets or rotations introduced during tissue sectioning and experimental handling, we need to apply an Affine Transformation to the image.
What is an Affine Transformation? An affine transformation is a linear mapping from 2D coordinates to 2D coordinates. It preserves the "collinearity" and "parallelism" of the image (i.e., straight lines remain straight, and parallel lines remain parallel). In the spatial transcriptomics registration process, we primarily leverage three geometric properties of affine transformations:
- Translation: Corrects the absolute positional shifts of the tissue section along the X and Y axes.
- Scaling: Corrects image size discrepancies caused by optical lens variations or spatial resolution differences.
- Rotation: Corrects the angular tilt introduced when the tissue section was mounted onto the spatial capture slide.
Core Transformation Parameters
In the subsequent steps, we aim to define a
Sx, Sy(Scale): The scaling factors of the image along the X/Y axes.Tx, Ty(Translate): The translation of the image along the X/Y axes, measured in original pixel distances.degree(Rotate): The rotation angle of the image around its center point.
Workflow Overview
To circumvent the trial-and-error approach of blindly guessing these parameters in the code, this tutorial provides an interactive solution:
- Automatically generate a local interactive HTML interface via code;
- Intuitively align the histological image with the transcriptomic cells through visual clicks within the interface;
- The web interface will automatically aggregate all your interactive adjustments into a standard affine matrix in the background using matrix multiplication. You simply need to copy the resulting
Sx, Sy, Tx, Ty, degreeparameters and apply them to the final Python OpenCV image processing function to achieve precise, full-image calibration.
Visualize the pre-registration status of the histological image
plt.figure(figsize=(20, 12))
# Plot the background histological image first
ax = plt.gca()
ax.imshow(img,
cmap='gray',
extent=[0, CHIP_MAX_X * RESOLUTION_UM, 0, CHIP_MAX_Y * RESOLUTION_UM], # Key: Define the image coordinate extent
aspect='auto',
alpha=1) # Set opacity level
# Plot the transcriptomic cell scatter overlay
ax.scatter(
table['X_um'],
table['Y_um'],
s=5,
alpha=0.8,
edgecolors='none',
rasterized=True
)
ax.set_aspect('equal')
Interactive Registration Tool
Since the Jupyter Notebook environment is prone to lag and rendering artifacts when handling massive scatter plots and high-frequency image redraws, we have adopted a more robust approach to ensure registration accuracy and fluidity: utilizing Python code to automatically generate a standalone, offline HTML interactive tool.
Technical Implementation and Principles:
- Offline Plotly.js: The script first extracts the
plotly.jslibrary from the local environment, eliminating loading delays caused by network latency. - Intelligent Image Downsampling: To guarantee ultra-fast browser rendering, the code proportionally downsamples the original high-resolution histological image in the background and converts it into a Base64 string embedded in the HTML. Note: This step only alters the visual display resolution; the backend matrix calculations remain strictly anchored to the image's original physical pixel dimensions.
User Instructions:
- Generate the Tool: Execute the code blocks below sequentially. Upon completion, a newly generated
alignment_tool.htmlfile will appear in your current working directory. - Open the Interface: Locate the HTML file in your system's file manager and double-click to open it in a web browser (Chrome or Edge is recommended).
- Interactive Fine-Tuning: Adjust the step size for each transformation in the top control panel. Click the translation, scaling, and rotation buttons to visually optimize the concordance between the histological background and the red scatter points (representing transcriptomic cells).
- Extract Parameters: The backend dynamically aggregates your operations into a standard 3x3 affine transformation matrix. Once you confirm the alignment is satisfactory, click the green "📋 Copy Parameters (JSON)" button.
- Apply Parameters: Paste the copied JSON parameters into the designated code cell in the subsequent sections of this tutorial to execute the final spatial transformation and cropping on the original high-resolution image.
# Extract pure local Plotly JS library
js_path = os.path.abspath("plotly_offline.min.js") # Replace with the actual path to the plotly JS file
with open(js_path, "w", encoding="utf-8") as f:
f.write(plotly.offline.get_plotlyjs())# Downsample histological image and convert to Base64 format
h_ori, w_ori = img_array.shape[:2]
max_dim = 1200
if max(h_ori, w_ori) > max_dim:
scale_down = max_dim / max(h_ori, w_ori)
img_small = cv2.resize(img_array, (int(w_ori * scale_down), int(h_ori * scale_down)))
else:
img_small = img_array
img_pil = Image.fromarray(img_small)
buffer = io.BytesIO()
img_pil.save(buffer, format="JPEG", quality=85)
img_b64 = "data:image/jpeg;base64," + base64.b64encode(buffer.getvalue()).decode('utf-8')# Subsample cell coordinates for rendering efficiency
sample_size = 80000
if len(table) > sample_size:
table_sample = table.sample(sample_size)
else:
table_sample = table
# Spatial cell coordinates mapping
# Parameters if using true physical distances (μm)
x_coords = [round(x, 2) for x in table_sample['X_um'].tolist()]
y_coords = [round(y, 2) for y in table_sample['Y_um'].tolist()]
W = float(CHIP_MAX_X * RESOLUTION_UM)
H = float(CHIP_MAX_Y * RESOLUTION_UM)
# Parameters if using raw chip grid coordinates
# x_coords = [round(x, 2) for x in table_sample['X'].tolist()]
# y_coords = [round(y, 2) for y in table_sample['Y'].tolist()]
# W = CHIP_MAX_X
# H = CHIP_MAX_Y
# Parameters if using optical image pixel coordinates
# x_coords = [round(x, 2) for x in table_sample['X_img'].tolist()]
# y_coords = [round(y, 2) for y in table_sample['Y_img'].tolist()]
# W = img_array.shape[1]
# H = img_array.shape[0]# ==========================================
# 3. Construct HTML (Corrected Scale Factor logic in affine matrix and added JSON copy feature)
# ==========================================
html_content = f"""
<!DOCTYPE html>
<html>
<head>
<meta charset="utf-8">
<title>SeekSpace Ultra-Fast Registration Tool</title>
<script src="./plotly_offline.min.js"></script>
<style> body {{ font-family: sans-serif; margin: 20px; background: #f8f9fa; }}
.panel {{ display: flex; gap: 15px; align-items: center; background: white; padding: 15px; border-radius: 8px; box-shadow: 0 2px 8px rgba(0,0,0,0.1); margin-bottom: 15px; flex-wrap: wrap; }}
button {{ padding: 6px 10px; cursor: pointer; border: 1px solid #ced4da; border-radius: 6px; background: #fff; font-size: 13px; transition: background 0.2s; }}
button:hover {{ background: #e9ecef; }}
input[type="number"] {{ width: 60px; padding: 4px; border: 1px solid #ced4da; border-radius: 4px; }}
.param-box {{ background: #212529; color: #61afef; padding: 10px 15px; border-radius: 6px; font-family: monospace; font-size: 14px; font-weight: bold; letter-spacing: 1px; }}
#plotly-div {{ width: 100%; height: 80vh; background: white; border-radius: 8px; box-shadow: 0 2px 8px rgba(0,0,0,0.1); }}
table { border: 0; }</style>
</head>
<body>
<div class="panel">
<div>
<b>Translate (<input type="number" id="step-val" value="30"> μm):</b>
<button onclick="moveImg('left')">⬅️ Left</button>
<button onclick="moveImg('right')">➡️ Right</button>
<button onclick="moveImg('up')">⬆️ Up</button>
<button onclick="moveImg('down')">⬇️ Down</button>
</div>
<div>
<b>Scale (<input type="number" id="step-s" value="0.01" step="0.01">):</b>
<button onclick="scaleImg('shrink')">➖ Shrink</button>
<button onclick="scaleImg('enlarge')">➕ Enlarge</button>
</div>
<div>
<b>Rotate (<input type="number" id="step-deg" value="0.25" step="0.25"> °):</b>
<button onclick="rotateImg(-1)">🔄 Counter-Clockwise</button>
<button onclick="rotateImg(1)">🔃 Clockwise</button>
</div>
<!-- Add right-aligned copy button and parameter display panel -->
<div style="margin-left: auto; display: flex; gap: 10px; align-items: center;">
<button onclick="copyParamsToClipboard()" style="background: #4caf50; color: white; border: none; font-weight: bold;">📋 Copy Parameters (JSON)</button>
<div class="param-box" id="param-display">Loading...</div>
</div>
</div>
<div id="plotly-div"></div>
<script>
const x_coords = {json.dumps(x_coords)};
const y_coords = {json.dumps(y_coords)};
const original_img_b64 = "{img_b64}";
const W = {W};
const H = {H};
const scale_xy = H / W;
// 1. Variable definitions (ported from Vue)
let pos_x = 0;
let pos_y = H;
let degree = 0;
let img_size_x = W;
let img_size_y = H;
// Core: Pure affine transformation matrix
let affine_matrix = [
[1.0, 0.0, 0.0],
[0.0, 1.0, 0.0],
[0.0, 0.0, 1.0]
];
// Matrix left-multiplication function (mimicking mathjs.multiply)
function multiplyMatrix(A, B) {{
let C = [[0,0,0],[0,0,0],[0,0,0]];
for(let i=0; i<3; i++)
for(let j=0; j<3; j++)
for(let k=0; k<3; k++)
C[i][j] += A[i][k] * B[k][j];
return C;
}}
function updateDisplay() {{
let Sx = affine_matrix[0][0];
let Sy = affine_matrix[1][1];
let Tx = affine_matrix[0][2];
let Ty = affine_matrix[1][2];
document.getElementById('param-display').innerText =
`Sx: ${{Sx.toFixed(4)}} | Sy: ${{Sy.toFixed(4)}} | Tx: ${{Tx.toFixed(2)}} | Ty: ${{Ty.toFixed(2)}} | Deg: ${{degree.toFixed(2)}}°`;
}}
// Function: Copy parameters to clipboard
window.copyParamsToClipboard = function() {{
let Sx = affine_matrix[0][0];
let Sy = affine_matrix[1][1];
let Tx = affine_matrix[0][2];
let Ty = affine_matrix[1][2];
// Construct standard JSON payload
let params = {{
"Sx": Number(Sx.toFixed(4)),
"Sy": Number(Sy.toFixed(4)),
"Tx": Number(Tx.toFixed(2)),
"Ty": Number(Ty.toFixed(2)),
"degree": Number(degree.toFixed(2))
}};
// Convert to formatted JSON string
let jsonStr = JSON.stringify(params, null, 2);
// Write to clipboard and trigger alert prompt
navigator.clipboard.writeText(jsonStr).then(() => {{
alert("✅ Affine transformation parameters successfully copied to clipboard!\\n\\n" + jsonStr);
}}).catch(err => {{
alert("❌ Copy failed, please manually copy the following:\\n" + jsonStr);
}});
}};
// 2. Translation logic fix: incorporate scale factor for accurate pixel mapping
window.moveImg = function(dir) {{
let s = parseFloat(document.getElementById('step-val').value) || 30;
let M = [[1, 0, 0], [0, 1, 0], [0, 0, 1]];
let current_scale = img_size_x / W;
let matrix_shift = s / current_scale;
if (dir === 'left') {{
M[0][2] = -matrix_shift;
pos_x -= s;
}} else if (dir === 'right') {{
M[0][2] = matrix_shift;
pos_x += s;
}} else if (dir === 'up') {{
M[1][2] = -matrix_shift;
pos_y += s;
}} else if (dir === 'down') {{
M[1][2] = matrix_shift;
pos_y -= s;
}}
affine_matrix = multiplyMatrix(M, affine_matrix);
Plotly.relayout('plotly-div', {{
'images[0].x': pos_x,
'images[0].y': pos_y
}});
updateDisplay();
}};
window.scaleImg = function(dir) {{
let s = parseFloat(document.getElementById('step-s').value) || 0.01;
let scale_factor = 1.0;
let delta_size = W * s;
if (dir === 'enlarge') {{
scale_factor = (img_size_x + delta_size) / img_size_x;
img_size_x += delta_size;
img_size_y += delta_size * scale_xy;
}} else if (dir === 'shrink') {{
scale_factor = (img_size_x - delta_size) / img_size_x;
img_size_x -= delta_size;
img_size_y -= delta_size * scale_xy;
}}
let M = [
[scale_factor, 0, 0],
[0, scale_factor, 0],
[0, 0, 1]
];
affine_matrix = multiplyMatrix(M, affine_matrix);
Plotly.relayout('plotly-div', {{
'images[0].sizex': img_size_x,
'images[0].sizey': img_size_y
}});
updateDisplay();
}};
window.rotateImg = function(sign) {{
const step = parseFloat(document.getElementById('step-deg').value) || 0.25;
degree += sign * step;
rotateBase64Image(original_img_b64, degree).then(rotatedB64 => {{
Plotly.relayout('plotly-div', {{'images[0].source': rotatedB64}});
}});
updateDisplay();
}};
function rotateBase64Image(base64data, degrees) {{
return new Promise((resolve, reject) => {{
const img = new Image();
img.src = base64data;
img.onload = function () {{
const canvas = document.createElement('canvas');
const ctx = canvas.getContext('2d');
canvas.width = img.width;
canvas.height = img.height;
ctx.translate(img.width / 2, img.height / 2);
ctx.rotate(degrees * (Math.PI / 180));
ctx.drawImage(img, -img.width / 2, -img.height / 2, img.width, img.height);
resolve(canvas.toDataURL());
}};
img.onerror = reject;
}});
}}
// Initial Plotly rendering
const data = [{{
x: x_coords,
y: y_coords,
mode: 'markers',
type: 'scattergl',
marker: {{ size: 2, color: 'red', opacity: 0.5 }},
name: 'Cells',
hoverinfo: 'none'
}}];
const layout = {{
xaxis: {{ range: [0, W], title: "X (um)", scaleanchor: "y", scaleratio: 1, showgrid: false }},
yaxis: {{ range: [0, H], title: "Y (um)", showgrid: false }},
margin: {{ l: 50, r: 50, t: 50, b: 50 }},
hovermode: false,
plot_bgcolor: 'black',
images: [{{
source: original_img_b64,
xref: "x",
yref: "y",
x: pos_x,
y: pos_y,
sizex: img_size_x,
sizey: img_size_y,
sizing: "stretch",
opacity: 0.6,
layer: "below"
}}]
}};
Plotly.newPlot('plotly-div', data, layout, {{responsive: true, displayModeBar: true}}).then(() => {{
updateDisplay();
}});
</script>
</body>
</html>
"""
html_filename = "alignment_tool.html"
html_path = os.path.abspath(html_filename)
with open(html_path, "w", encoding="utf-8") as f:
f.write(html_content)
print(f"✅ Interactive HTML registration tool generated successfully. Please double-click to open: {html_filename}")Apply Affine Transformation to the Histological Image
Please paste the affine transformation parameters derived from the interactive registration tool into the params dictionary below. It must contain the following 5 keys: Sx, Sy, Tx, Ty, degree.
def affine_transformation(img, Sx, Sy, Tx, Ty, degree):
h, w, c = img.shape
x0 = w/2
y0 = h/2
M_rotate = np.array([[np.cos(np.radians(degree)), -np.sin(np.radians(degree)), x0 * (1 - np.cos(np.radians(degree))) + y0 * np.sin(np.radians(degree))],
[np.sin(np.radians(degree)), np.cos(np.radians(degree)), y0 * (1 - np.cos(np.radians(degree))) - x0 * np.sin(np.radians(degree))],
[0, 0, 1]])
M_trans = np.array([[Sx, 0, Tx],
[0, Sy, Ty],
[0, 0, 1]])
M_f = np.dot(M_trans, M_rotate)
# cv2.warpAffine provides strict affine transformation (better computational performance than warpPerspective)
# img_transformed = cv2.warpAffine(img, M_trans, (w, h), borderValue=(0, 0, 0))
return cv2.warpPerspective(img, M_f, (w, h), borderValue=(255, 255, 255))
params = {
"Sx": 0.994,
"Sy": 0.994,
"Tx": 79.86,
"Ty": 39.46,
"degree": -0.45
}
img_trans = affine_transformation(img_array, **params)Visualize Registration Concordance (Transformed Image vs. Cells)
Finally, we overlay the spatially transformed image (img_trans) with the transcriptomic cell scatter coordinates to visually validate the registration accuracy.
⚠️ Crucial Note: Limitations of Rigid Transformations
Please be aware that the methodology provided in this tutorial utilizes Affine Transformation. Affine transformations fall under the category of rigid/semi-rigid transformations, which solely account for global translation, rotation, and scaling. It inherently assumes the entire tissue section acts as a rigid body, where the relative spatial topologies between internal cellular structures remain absolutely constant.
However, in authentic biological experiments—particularly when aligning adjacent consecutive sections or working with large-area tissue mounts—tissues frequently undergo localized Non-rigid Deformations (Elastic Deformations) such as tearing, folding, or warping.
If your histological image exhibits such localized non-rigid deformations, achieving perfect cell-to-morphology alignment is impossible relying exclusively on the global rigid affine transformations presented in this tutorial.
In such scenarios, we strongly recommend the following protocol: Prior to executing downstream spatial transcriptomic coordinate alignment, import your raw histological images into specialized image registration software to perform Elastic/Non-linear Registration. Industry-standard tools for this include:
- Fiji/ImageJ (e.g., utilizing plugins like TrakEM2 or bUnwarpJ)
- QuPath (for whole-slide image analysis and registration)
- Standalone algorithmic registration frameworks such as Elastix.
plt.figure(figsize=(20, 12))
# Plot the background histological image first
ax = plt.gca()
ax.imshow(img_trans,
cmap='gray',
extent=[0, CHIP_MAX_X * RESOLUTION_UM, 0, CHIP_MAX_Y * RESOLUTION_UM], # Key: Define the image coordinate extent
aspect='auto',
alpha=1) # Set opacity level
# Plot the transcriptomic cell scatter overlay
ax.scatter(
table['X_um'],
table['Y_um'],
s=5,
alpha=0.8,
edgecolors='none',
rasterized=True
)
ax.set_aspect('equal')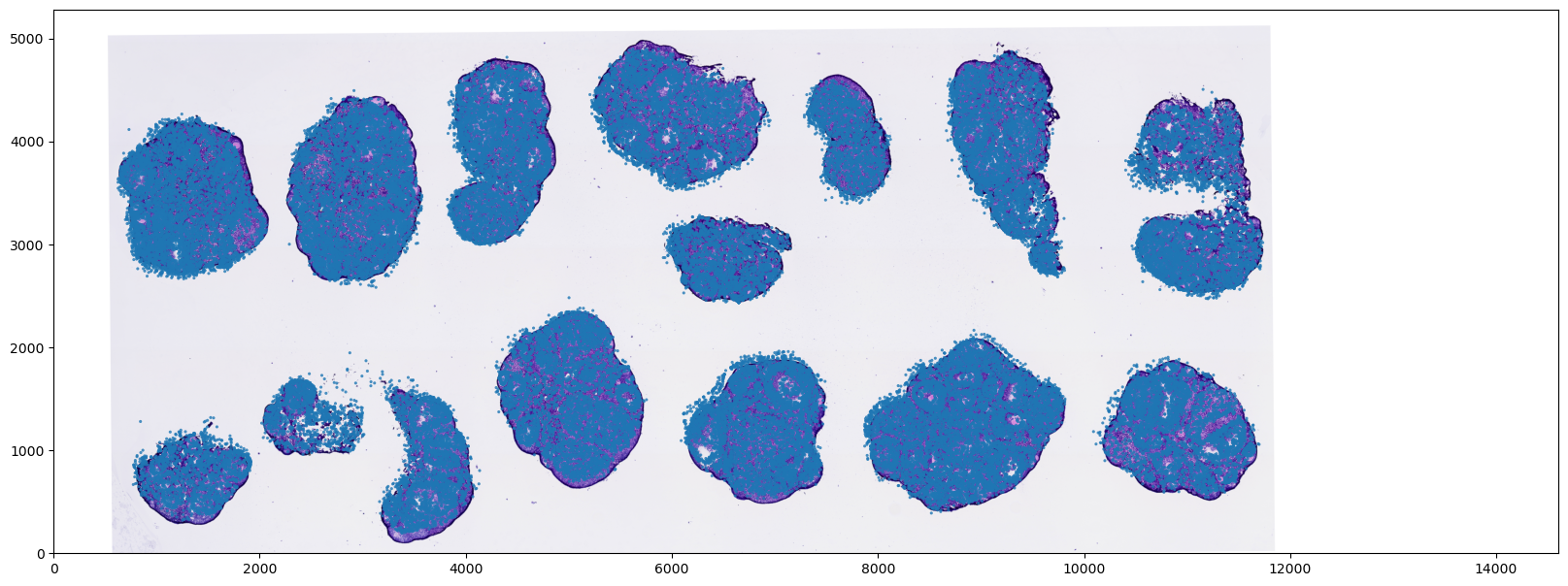
Save the spatially registered histological image
output_filename = "aligned_HE_transformed.png"
final_img = Image.fromarray(img_trans.astype('uint8'))
final_img.save(output_filename)