Single-cell Spatial Neighborhood Statistics: Spatial Distribution and Neighborhood Analysis
Time: 12 min
Words: 2.4k words
Updated: 2026-02-28
Reads: 0 times
Load Analysis R Packages
R
library(Seurat)
library(qs)
library(dplyr)
library(tibble)
library(stringr)
library(ggplot2)
library(phenoptr)output
Attaching SeuratObject
qs 0.27.3. Announcement: https://github.com/qsbase/qs/issues/103
Warning message:
“package ‘dplyr’ was built under R version 4.3.3”
Attaching package: ‘dplyr’
The following objects are masked from ‘package:stats’:
filter, lag
The following objects are masked from ‘package:base’:
intersect, setdiff, setequal, union
Warning message:
“package ‘tibble’ was built under R version 4.3.3”
Warning message:
“package ‘stringr’ was built under R version 4.3.3”
Warning message:
“package ‘ggplot2’ was built under R version 4.3.3”
qs 0.27.3. Announcement: https://github.com/qsbase/qs/issues/103
Warning message:
“package ‘dplyr’ was built under R version 4.3.3”
Attaching package: ‘dplyr’
The following objects are masked from ‘package:stats’:
filter, lag
The following objects are masked from ‘package:base’:
intersect, setdiff, setequal, union
Warning message:
“package ‘tibble’ was built under R version 4.3.3”
Warning message:
“package ‘stringr’ was built under R version 4.3.3”
Warning message:
“package ‘ggplot2’ was built under R version 4.3.3”
Input Object Details
- col_SAM: Sample column name in metadata
- SAMple_name: Sample name to analyze
- col_celltype: Cell type column name in metadata
- analysis_celltype: Cell type to analyze
R
col_sam = "Sample"
sample_name = "T1"
col_celltype = "large_celltype"
analysis_celltype = "Neutrophils"Read RDS and Metadata, Preprocess Data
R
data = readRDS("../../data/AY1739779861133/input.rds")R
meta = read.delim("../../data/AY1739779861133/meta.tsv")
rownames(meta) = meta$barcode
data@meta.data = meta- Extract spatial sample info to analyze from RDS
- Convert SeekSpace spatial coordinates: Pixel -> Actual (Conversion factor 0.2653)
- Extract cell types to analyze
- Calculate distances between each cell point
R
sub = subset(data,cells = rownames(data@meta.data[data@meta.data[[col_sam]] %in% sample_name,]))
sub@meta.data$barcode = rownames(sub@meta.data)
df_labels = sub@meta.data[,c("barcode",col_celltype)]
sub_position = Embeddings(sub,"spatial")
sub_position = sub_position*0.2653
sub_position = as.data.frame(sub_position)
Neu_barcode = rownames(sub@meta.data[sub@meta.data[[col_celltype]] %in% analysis_celltype,])
dist_matrix <- as.matrix(dist(sub_position, method = "euclidean"))Calculate Spatial Neighborhood Composition in 4 Ranges
R
neigh_freq1 = data.frame()
for(threshold in c(20,50,100,200)){
# Find neighbors for each point (excluding self)
neighbors = apply(dist_matrix, 1, function(row) {
nearby = which(row <= threshold & row > 0) # row > 0 excludes self
if (length(nearby) > 0) {
paste(names(nearby), collapse = ";") # Semicolon separated neighbors
} else {
NA # Return NA if no neighbors
}
})
# Convert result to data frame
result = data.frame(
Point = rownames(sub_position),
X = sub_position$spatial_1,
Y = sub_position$spatial_2,
Neighbors = neighbors,
stringsAsFactors = FALSE
)
result_Neu = result[result$Point %in% Neu_barcode,]
map_neighbors_to_labels = function(neighbor_ids, label_df) {
if (is.na(neighbor_ids)) return(NA)
ids = strsplit(neighbor_ids, ";")[[1]] # Split IDs
labels = label_df$large_celltype[match(ids, label_df$barcode)] # Match cell types
paste(labels, collapse = "; ") # Merge to string
}
# Apply to Neighbors column
result_Neu$Neighbor_Labels = sapply(
result_Neu$Neighbors,
map_neighbors_to_labels,
label_df = df_labels
)
merged_string = paste(result_Neu$Neighbors, collapse = ";")
a = str_split(merged_string,";")[[1]]
neigh = data.frame(barcode = unique(a))
neigh = left_join(neigh,sub@meta.data[,c("barcode",col_celltype)])
neigh_freq = as.data.frame(table(neigh[[col_celltype]]))
colnames(neigh_freq) = c("Celltype","Cell_number")
neigh_freq$Distance = paste0(threshold,"um")
neigh_freq1 = rbind(neigh_freq1,neigh_freq)
}output
Joining with \`by = join_by(barcode)\`
Joining with \`by = join_by(barcode)\`
Joining with \`by = join_by(barcode)\`
Joining with \`by = join_by(barcode)\`
Joining with \`by = join_by(barcode)\`
Joining with \`by = join_by(barcode)\`
Joining with \`by = join_by(barcode)\`
R
neigh_freq1| Celltype | Cell_number | Distance |
|---|---|---|
| <fct> | <int> | <chr> |
| B_cells | 49 | 20um |
| Endothelial_cells | 9 | 20um |
| Epithelial_cells | 68 | 20um |
| Fibroblasts | 105 | 20um |
| Macrophages | 35 | 20um |
| Mast_cells | 2 | 20um |
| Nerve | 3 | 20um |
| Neutrophils | 39 | 20um |
| Plasma_cells | 34 | 20um |
| SMC | 4 | 20um |
| T_cells | 94 | 20um |
| B_cells | 188 | 50um |
| Endothelial_cells | 41 | 50um |
| Epithelial_cells | 443 | 50um |
| Fibroblasts | 312 | 50um |
| Macrophages | 106 | 50um |
| Mast_cells | 4 | 50um |
| Nerve | 7 | 50um |
| Neutrophils | 162 | 50um |
| Plasma_cells | 119 | 50um |
| SMC | 35 | 50um |
| T_cells | 310 | 50um |
| B_cells | 386 | 100um |
| Endothelial_cells | 82 | 100um |
| Epithelial_cells | 1401 | 100um |
| Fibroblasts | 596 | 100um |
| Macrophages | 188 | 100um |
| Mast_cells | 5 | 100um |
| Nerve | 10 | 100um |
| Neutrophils | 289 | 100um |
| Plasma_cells | 235 | 100um |
| SMC | 79 | 100um |
| T_cells | 556 | 100um |
| B_cells | 603 | 200um |
| Endothelial_cells | 111 | 200um |
| Epithelial_cells | 2902 | 200um |
| Fibroblasts | 800 | 200um |
| Macrophages | 262 | 200um |
| Mast_cells | 6 | 200um |
| Nerve | 14 | 200um |
| Neutrophils | 383 | 200um |
| Plasma_cells | 307 | 200um |
| SMC | 121 | 200um |
| T_cells | 750 | 200um |
- Calculate neighborhood cell type proportions around target cell type within each distance range
R
neigh_freq1 = neigh_freq1 %>%
group_by(Distance) %>% # Step 1: Group by Region
mutate(Region_Pct = Cell_number / sum(Cell_number) * 100) %>% # Step 2: Calculate within-group percentage
ungroup()
neigh_freq1$Distance = factor(neigh_freq1$Distance,c("20um","50um","100um","200um"))R
neigh_freq1| Celltype | Cell_number | Distance | Region_Pct |
|---|---|---|---|
| <fct> | <int> | <fct> | <dbl> |
| B_cells | 49 | 20um | 11.08597285 |
| Endothelial_cells | 9 | 20um | 2.03619910 |
| Epithelial_cells | 68 | 20um | 15.38461538 |
| Fibroblasts | 105 | 20um | 23.75565611 |
| Macrophages | 35 | 20um | 7.91855204 |
| Mast_cells | 2 | 20um | 0.45248869 |
| Nerve | 3 | 20um | 0.67873303 |
| Neutrophils | 39 | 20um | 8.82352941 |
| Plasma_cells | 34 | 20um | 7.69230769 |
| SMC | 4 | 20um | 0.90497738 |
| T_cells | 94 | 20um | 21.26696833 |
| B_cells | 188 | 50um | 10.88592936 |
| Endothelial_cells | 41 | 50um | 2.37405906 |
| Epithelial_cells | 443 | 50um | 25.65141865 |
| Fibroblasts | 312 | 50um | 18.06601042 |
| Macrophages | 106 | 50um | 6.13781123 |
| Mast_cells | 4 | 50um | 0.23161552 |
| Nerve | 7 | 50um | 0.40532716 |
| Neutrophils | 162 | 50um | 9.38042849 |
| Plasma_cells | 119 | 50um | 6.89056167 |
| SMC | 35 | 50um | 2.02663578 |
| T_cells | 310 | 50um | 17.95020266 |
| B_cells | 386 | 100um | 10.08622942 |
| Endothelial_cells | 82 | 100um | 2.14267050 |
| Epithelial_cells | 1401 | 100um | 36.60830938 |
| Fibroblasts | 596 | 100um | 15.57355631 |
| Macrophages | 188 | 100um | 4.91246407 |
| Mast_cells | 5 | 100um | 0.13065064 |
| Nerve | 10 | 100um | 0.26130128 |
| Neutrophils | 289 | 100um | 7.55160700 |
| Plasma_cells | 235 | 100um | 6.14058009 |
| SMC | 79 | 100um | 2.06428011 |
| T_cells | 556 | 100um | 14.52835119 |
| B_cells | 603 | 200um | 9.63412686 |
| Endothelial_cells | 111 | 200um | 1.77344624 |
| Epithelial_cells | 2902 | 200um | 46.36523406 |
| Fibroblasts | 800 | 200um | 12.78159450 |
| Macrophages | 262 | 200um | 4.18597220 |
| Mast_cells | 6 | 200um | 0.09586196 |
| Nerve | 14 | 200um | 0.22367790 |
| Neutrophils | 383 | 200um | 6.11918837 |
| Plasma_cells | 307 | 200um | 4.90493689 |
| SMC | 121 | 200um | 1.93321617 |
| T_cells | 750 | 200um | 11.98274485 |
Result Visualization
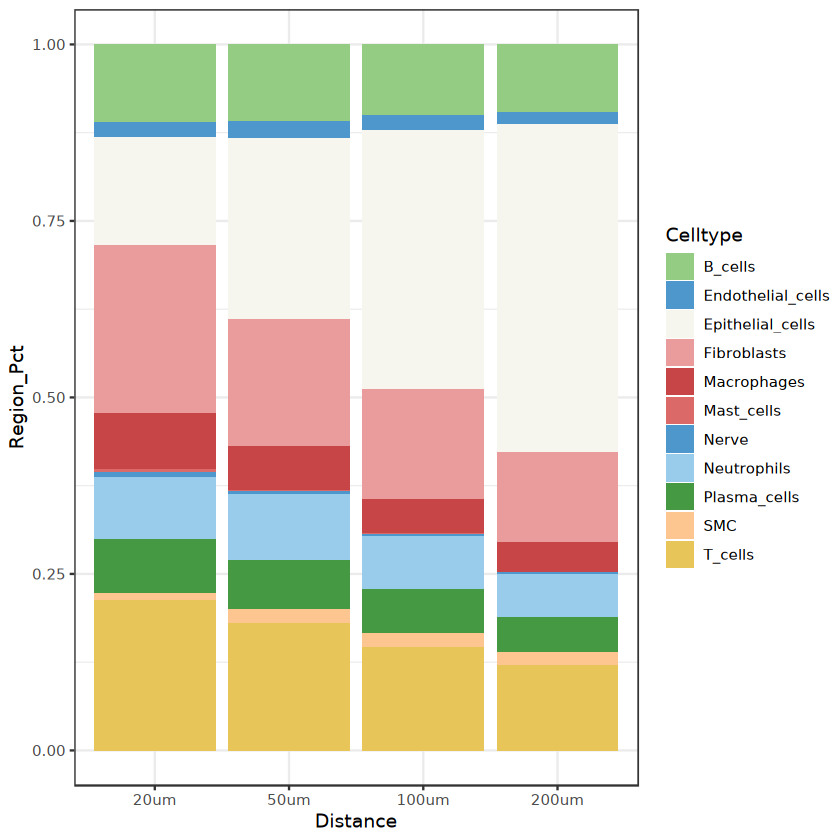
Visualize proportions of surrounding cell types around target cell type
R
ggplot(neigh_freq1, aes(fill = Celltype, y = Region_Pct, x = Distance)) +
theme_bw() +
geom_bar(position = "fill", stat = "identity") +
scale_fill_manual(
values = c("#93cc82", "#4d97cd", "#f6f5ee", "#ea9c9d", "#c74546",
"#db6968", "#4d97cd", "#99cbeb", "#459943",
"#fdc58f", "#e8c559", "#a3d393", "#f8984e"), # Customize colors
name = "Celltype" # Custom legend name
) +
theme(
plot.title = element_text(hjust = 0.5),
panel.background = element_blank()
)
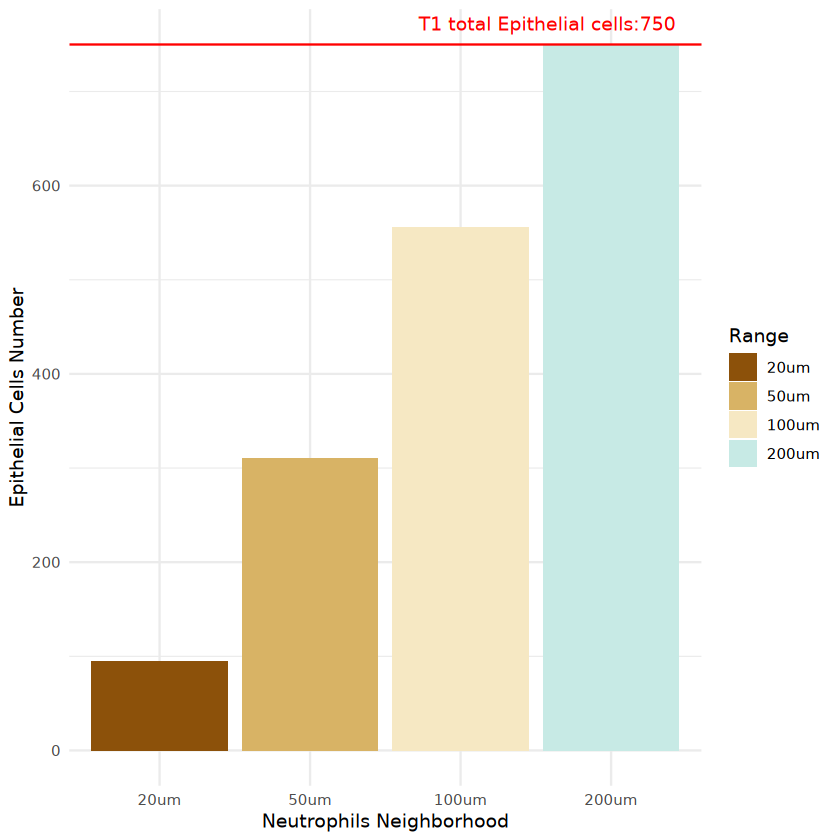
Visualize counts of specific surrounding cell types around target cell type
R
plot = neigh_freq1[neigh_freq1$Celltype %in% "T_cells",]
plot$Distance = factor(plot$Distance,levels = c("20um","50um","100um","200um"))R
plot| Celltype | Cell_number | Distance | Region_Pct |
|---|---|---|---|
| <fct> | <int> | <fct> | <dbl> |
| T_cells | 94 | 20um | 21.26697 |
| T_cells | 310 | 50um | 17.95020 |
| T_cells | 556 | 100um | 14.52835 |
| T_cells | 750 | 200um | 11.98274 |
R
ggplot(plot, aes(x = Distance, y = Cell_number, fill = Distance)) +
geom_col() + # Use geom_col() instead of geom_bar(stat="identity")
scale_fill_manual(
values = c("#8c510a", "#d8b365", "#f6e8c3", "#c7eae5"),
name = "Range" # Custom legend name
) +
labs(
x = "Neutrophils Neighborhood", # x-axis label
y = "Epithelial Cells Number", # y-axis label
) +
theme_minimal()+ # Use minimal theme
geom_hline(yintercept = 750,color="red")+
annotate(
"text",
x = Inf, # Display on far right
y = 750,
label = "T1 total Epithelial cells:750",
color = "red",
hjust = 1.1, # Right align
vjust = -1
)
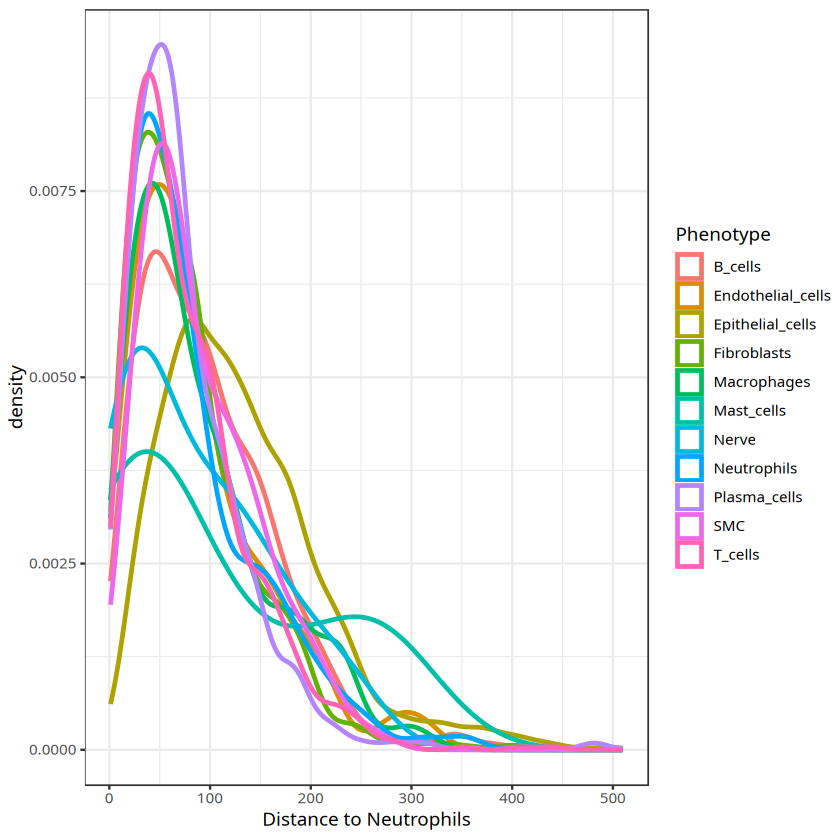
Visualize density distribution of cell types around target cell type
R
sub_position = rownames_to_column(sub_position,"Cell ID")
sub_position = cbind(sub_position,sub@meta.data[,col_celltype])
colnames(sub_position)[2:4] = c("Cell X Position","Cell Y Position","Phenotype")
sub_positionR
distances=find_nearest_distance(sub_position)
csd_with_distance=bind_cols(sub_position, distances)
ggplot(csd_with_distance,aes(`Distance to Neutrophils`, color=Phenotype))+geom_density(size=1)+theme_bw()