SpaGene Advanced Spatial Transcriptomics Analysis: Spatially Variable Genes and Ligand-Receptor Colocalization
Background Introduction
Method Principle
Given a set of spatial locations, SpaGene first constructs a spatial network using k-nearest neighbors.
For each gene, SpaGene then extracts a subnetwork from the k-nearest neighbor graph, containing only cells/spots highly expressing that gene.
SpaGene quantifies subnetwork connectivity by calculating Earth Mover's Distance (EMD) between the subnetwork's degree distribution and that of a fully connected network.
Finally, SpaGene compares observed distances with expected distances obtained via random permutation.
Method Application
SpaGene identifies spatially variable genes and their distribution patterns by quantifying spatial clustering of gene expression.
Ligand-Receptor Colocalization: Reveals potential mechanisms of intercellular communication by analyzing spatial co-expression patterns of ligands and receptors.
Combining SpaGene and ligand-receptor colocalization analysis allows for more comprehensive study of biological significance in spatial transcriptomics data.
SpaGene Calculation Results
library(SpaGene)
library(Seurat)
library(pheatmap)
library(ggplot2)
library(stringr)
library(tibble)Loading required package: ggplot2
Loading required package: patchwork
Attaching SeuratObject
Warning message:
“package ‘stringr’ was built under R version 4.3.3”
Warning message:
“package ‘tibble’ was built under R version 4.3.3”
SpaGene Parameter Description
- rds_path: Path to rds
- meta_path: Path to meta
- col_SAM: Corresponding sample column name in meta, multiple samples separated by commas, e.g., "A,B,C"
- SAMple_name: Sample name, corresponding to values under col_SAM column
- species: Species selection, "human" or "mouse"
rds_path="../../data/AY1748480899609//input.rds"
meta_path="../../data/AY1748480899609//meta.tsv"
col_sam="Sample"
sample_name="N1_expression"
species="human"data = readRDS(rds_path)
meta <- read.table(meta_path,header=T,sep='\t',check.names=F)
if(colnames(meta)[1]=="barcode"){
rownames(meta) <- meta$barcode
} else if(colnames(meta)[1]=="barcodes"){
rownames(meta) <- meta$barcodes
}
data@meta.data <- meta
sample_name = str_split(sample_name,",")[[1]]
ll=list()
ll_heatmap = list()
for(i in 1:length(sample_name)){
data_subset = subset(data,cells = rownames(data@meta.data[data@meta.data[[col_sam]] %in% sample_name[i],]))
position = data_subset@reductions$spatial@cell.embeddings
data_subset_count = GetAssayData(data_subset,assay = "RNA",slot = "count")
spagene = SpaGene(data_subset_count,position)
pattern=FindPattern(spagene)
ll[[sample_name[i]]]$pattern = pattern
ll[[sample_name[i]]]$position = position
top5<-apply(pattern$genepattern,2,function(x){names(x)[order(x,decreasing=T)][1:5]})
ll_heatmap[[sample_name[i]]] = pheatmap(pattern$genepattern[rownames(pattern$genepattern)%in%top5,])
a = pattern$pattern
colnames(a) = rownames(position)
write.table(a,paste0(sample_name[i],"_pattern_expression.txt"),sep = "\t",col.names = T,row.names = T,quote = F)
}
Spatial Expression Patterns
options(repr.plot.height=8, repr.plot.width=16)
for(i in 1:length(sample_name)){
print(sample_name[i])
plot = PlotPattern(ll[[sample_name[i]]]$pattern,ll[[i]]$position,pt.size = 0.3)
print(plot)
}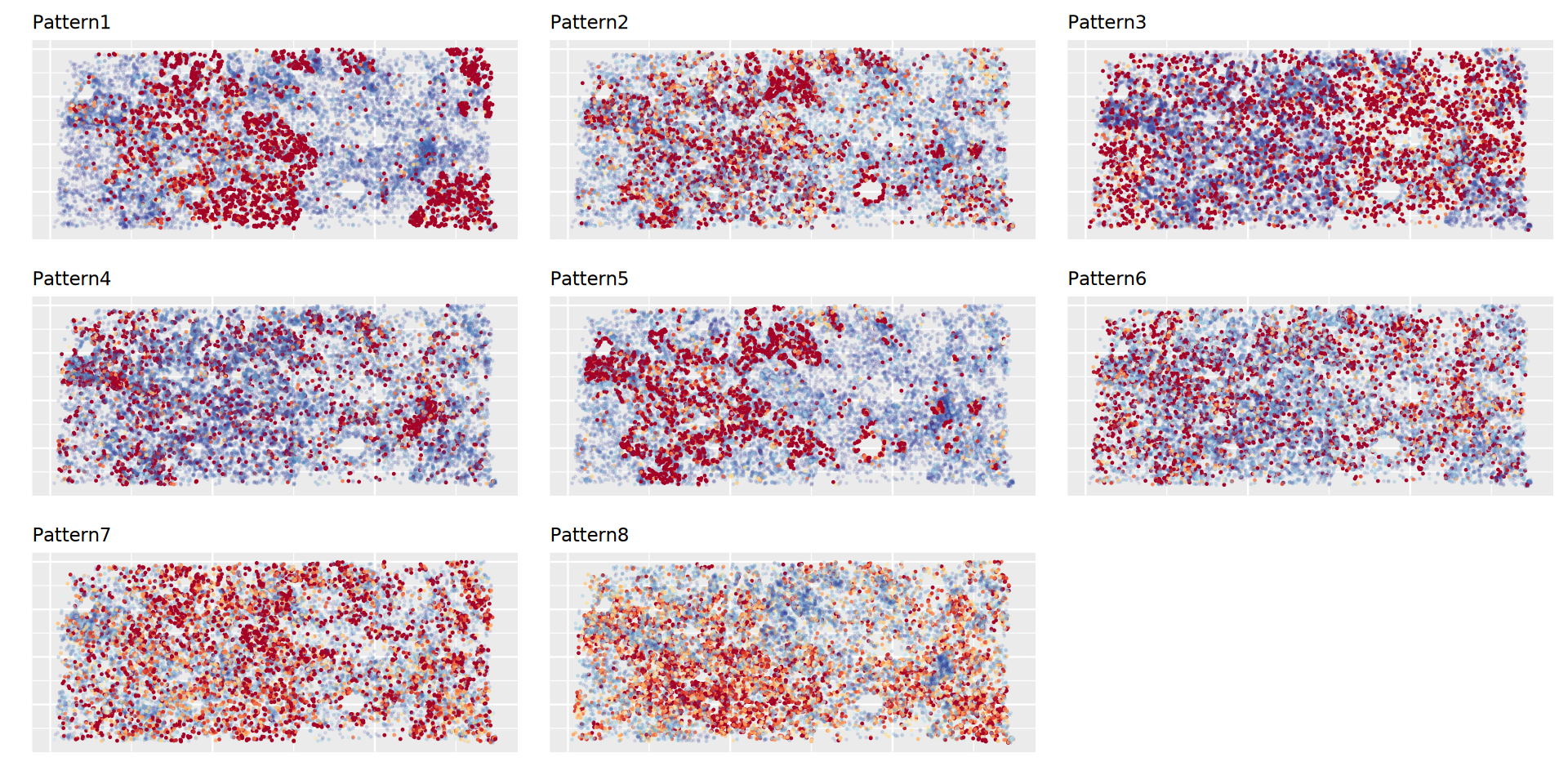
Figure shows spatial expression of gene patterns; darker red indicates higher expression, darker blue indicates lower expression.Gene Expression Heatmap
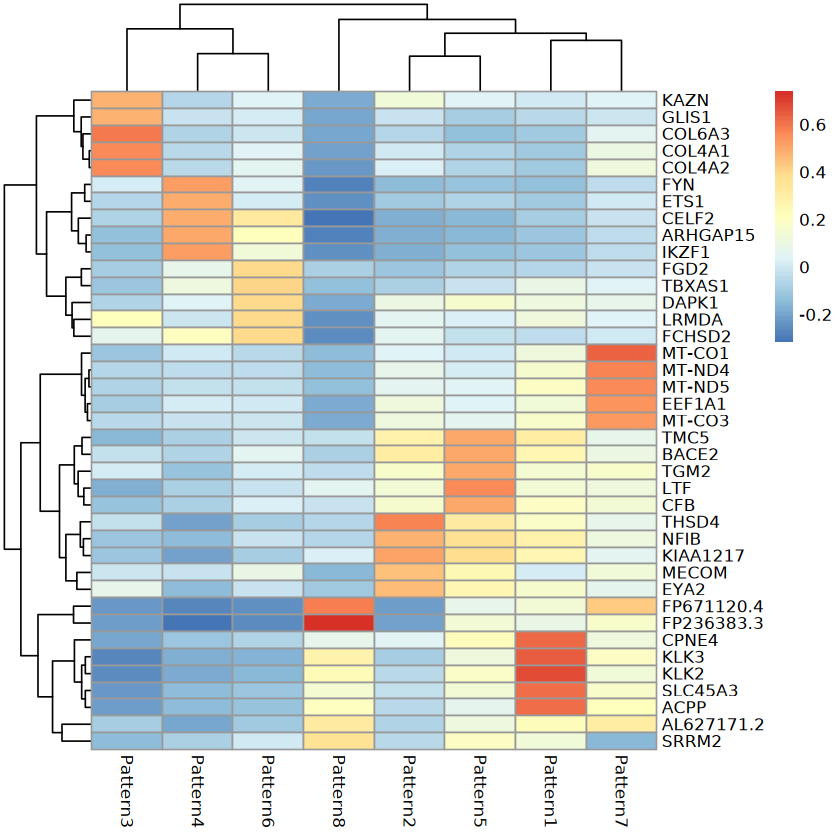
Expression Heatmap of Top 5 Genes in Each Pattern
for(i in 1:length(sample_name)){
print(sample_name[i])
print(ll_heatmap[[sample_name[i]]])
}
Heatmap shows expression of Top 5 genes in each pattern; X-axis represents genes; Y-axis represents patterns.Ligand-Receptor Gene Spatial Expression
Load Ligand-Receptor Database, Optional Methods:
- Download ligand-receptor databases provided by common cell communication tools
- Ligand-Receptor database provided in SpaGene tool GitHub
- Your own library of interest
Input ligand-receptor database file requires two columns named (ligand_gene_symbol and receptor_gene_symbol), with row names as LigandGene_ReceptorGene
if(species == "human"){
LRpair = read.delim("human_LR.txt",sep = "\t",row.names = 1)
} else if(species == "mouse"){
LRpair = read.delim("mouse_LR.txt",sep = "\t",row.names = 1)
}ll_plotLR = list()
for(i in 1:length(sample_name)){
data_subset = subset(data,cells = rownames(data@meta.data[data@meta.data[[col_sam]] %in% sample_name[i],]))
position = data_subset@reductions$spatial@cell.embeddings
data_subset_count = GetAssayData(data_subset,assay = "RNA",slot = "count")
data_lr = SpaGene_LR(data_subset_count,position,LRpair=LRpair)
data_lr = data_lr[order(data_lr$adj),]
# You can select LR pairs to display co-expression using LRpair parameter
ll_plotLR[[sample_name[i]]] = plotLR(data_subset_count,position,LRpair=c(gsub(rownames(data_lr)[1],pattern = ".*[_]",replacement = ""),gsub(rownames(data_lr)[1],pattern = "[_].*",replacement = "")),alpha.min=0.2,pt.size = 0.3)
}“sparse->dense coercion: allocating vector of size 5.4 GiB”
Calculate significant ligand-receptor colocalization and co-expression pairs, visualizing spatial distribution of LR pairs
options(repr.plot.height=4, repr.plot.width=16)
for(i in 1:length(sample_name)){
print(sample_name[i])
print(ll_plotLR[[sample_name[i]]])
}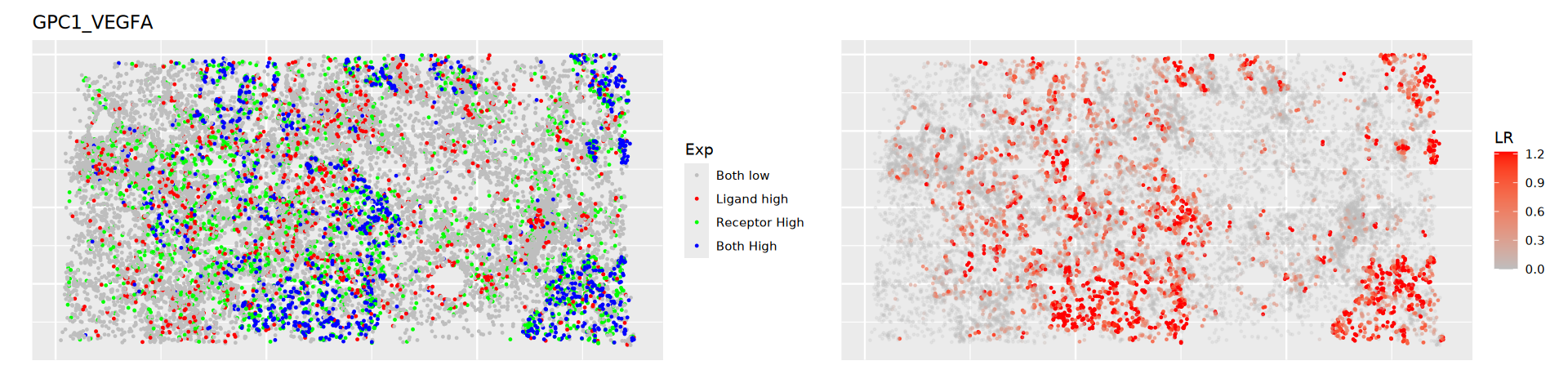
- Left Plot: Gray represents cells with low receptor and ligand expression; Red represents high ligand expression; Green represents high receptor expression; Blue represents high expression of both.
- Right Plot: Ligand-Receptor pair expression score; darker color indicates higher likelihood of interaction.Literature Case Analysis
Literature 1: Literature 'Single-cell and spatial transcriptome analyses reveal tertiary lymphoid structures linked to tumour progression and immunotherapy response in nasopharyngeal carcinoma': Previous cell communication analysis found CXCL13+ CAFs interact with B cells via CXCL13-CXCR5, VCAM1-ITGAM2 pairs. SpaGene further confirmed these pairs are spatially proximal and co-expressed, verifying the interaction between CXCL13+ CAFs and B cells.
Literature 2: Literature 'A Specialized Epithelial Cell Type Regulating Mucosal Immunity and Driving Human Crohn’s Disease': To explore interactions between epithelial and immune cells, SpaGene was used to find that LND-high marker genes were significantly correlated with high-expression immune cell signaling genes and spatially proximal.
References
[1] Pang Z, Chong J, Zhou G, de Lima Morais DA, Chang L, Barrette M, et al. MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 2021;49:W388–96.
[2] Liu, Y., Ye, SY., He, S. et al. Single-cell and spatial transcriptome analyses reveal tertiary lymphoid structures linked to tumour progression and immunotherapy response in nasopharyngeal carcinoma. Nat Commun 15, 7713 (2024).
[3] Li J, Simmons AJ, Chiron S, Ramirez-Solano MA, Tasneem N, Kaur H, Xu Y, Revetta F, Vega PN, Bao S, Cui C, Tyree RN, Raber LW, Conner AN, Beaulieu DB, Dalal RL, Horst SN, Pabla BS, Huo Y, Landman BA, Roland JT, Scoville EA, Schwartz DA, Washington MK, Shyr Y, Wilson KT, Coburn LA, Lau KS, Liu Q. A Specialized Epithelial Cell Type Regulating Mucosal Immunity and Driving Human Crohn's Disease. bioRxiv [Preprint]. 2023 Oct 2:2023.09.30.560293. doi: 10.1101/2023.09.30.560293. Update in: Nat Commun. 2024 Aug 22;15(1):7204.
