SeekSpace 空间转录组多样本整合分析 (Seurat 框架)
时长: 17 分钟
字数: 3.3k 字
更新: 2026-02-28
阅读: 0 次
R
library(Seurat)
library(tidyverse)
library(ggplot2)
library(harmony)多样本整合
R
samples <- c("WTH1059","WTH1092")
path <- "./demo_integrated"R
obj.list <- list()
info2keep <- list()
numsap=1
for (each in samples){
obj <- readRDS(paste0(path,'/',each,'.rds'))
obj$stim <-each
obj.list[[each]] <- obj
#保留单样本rds中的图片信息
if (length(obj@misc$info) > 0) {
info2keep <- append(info2keep, obj@misc$info)
}
}R
sub.ob<-obj.list[-1]
#merge时添加merge.dr = "spatial",保留reductions中的spatial坐标
obj.integrated <- merge(obj.list[[1]], sub.ob, merge.dr = "spatial")
if (length(info2keep) > 0) { obj.integrated@misc$info <- info2keep }R
str(obj.integrated)output
Formal class 'Seurat' [package "SeuratObject"] with 13 slots
..@ assays :List of 1
.. ..$ RNA:Formal class 'Assay' [package "SeuratObject"] with 8 slots
.. .. .. ..@ counts :Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
.. .. .. .. .. ..@ i : int [1:47683687] 493 876 1115 1617 1838 1962 2169 2613 2959 4318 ...n .. .. .. .. .. ..@ p : int [1:59172] 0 117 197 271 402 469 520 669 754 890 ...n .. .. .. .. .. ..@ Dim : int [1:2] 32285 59171
.. .. .. .. .. ..@ Dimnames:List of 2
.. .. .. .. .. .. ..$ : chr [1:32285] "Xkr4" "Gm1992" "Gm19938" "Gm37381" ...n .. .. .. .. .. .. ..$ : chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n .. .. .. .. .. ..@ x : num [1:47683687] 1 1 1 1 1 1 1 1 1 1 ...n .. .. .. .. .. ..@ factors : list()
.. .. .. ..@ data :Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
.. .. .. .. .. ..@ i : int [1:47683687] 493 876 1115 1617 1838 1962 2169 2613 2959 4318 ...n .. .. .. .. .. ..@ p : int [1:59172] 0 117 197 271 402 469 520 669 754 890 ...n .. .. .. .. .. ..@ Dim : int [1:2] 32285 59171
.. .. .. .. .. ..@ Dimnames:List of 2
.. .. .. .. .. .. ..$ : chr [1:32285] "Xkr4" "Gm1992" "Gm19938" "Gm37381" ...n .. .. .. .. .. .. ..$ : chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n .. .. .. .. .. ..@ x : num [1:47683687] 3.93 3.93 3.93 3.93 3.93 ...n .. .. .. .. .. ..@ factors : list()
.. .. .. ..@ scale.data : num[0 , 0 ]
.. .. .. ..@ key : Named chr "rna_"
.. .. .. .. ..- attr(*, "names")= chr ""
.. .. .. ..@ assay.orig : NULL
.. .. .. ..@ var.features : logi(0)
.. .. .. ..@ meta.features:'data.frame': 32285 obs. of 0 variables
.. .. .. ..@ misc : list()
..@ meta.data :'data.frame': 59171 obs. of 12 variables:
.. ..$ orig.ident : chr [1:59171] "demo" "demo" "demo" "demo" ...n .. ..$ nCount_RNA : num [1:59171] 200 200 200 200 200 200 200 200 200 200 ...n .. ..$ nFeature_RNA : int [1:59171] 117 80 74 131 67 51 149 85 136 49 ...n .. ..$ percent.mito : num [1:59171] 0 0.5 1.5 0 0.5 0 0 0.5 0 0.5 ...n .. ..$ RNA_snn_res.0.2: chr [1:59171] "5" "6" "3" "1" ...n .. ..$ RNA_snn_res.0.5: chr [1:59171] "2" "3" "19" "12" ...n .. ..$ RNA_snn_res.0.8: chr [1:59171] "2" "16" "23" "15" ...n .. ..$ RNA_snn_res.1.1: chr [1:59171] "1" "7" "23" "17" ...n .. ..$ RNA_snn_res.1.4: chr [1:59171] "5" "2" "24" "17" ...n .. ..$ seurat_clusters: chr [1:59171] "5" "2" "24" "17" ...n .. ..$ Sample : chr [1:59171] "demo" "demo" "demo" "demo" ...n .. ..$ stim : chr [1:59171] "WTH1059" "WTH1059" "WTH1059" "WTH1059" ...n ..@ active.assay: chr "RNA"
..@ active.ident: Factor w/ 28 levels "0","1","10","11",..: 13 9 17 8 15 3 26 15 8 26 ...n .. ..- attr(*, "names")= chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n ..@ graphs : list()
..@ neighbors : list()
..@ reductions :List of 1
.. ..$ spatial:Formal class 'DimReduc' [package "SeuratObject"] with 9 slots
.. .. .. ..@ cell.embeddings : int [1:59171, 1:2] 34524 42576 44264 40223 36766 33959 26542 28047 39761 21306 ...n .. .. .. .. ..- attr(*, "dimnames")=List of 2
.. .. .. .. .. ..$ : chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n .. .. .. .. .. ..$ : chr [1:2] "spatial_1" "spatial_2"
.. .. .. ..@ feature.loadings : num[0 , 0 ]
.. .. .. ..@ feature.loadings.projected: num[0 , 0 ]
.. .. .. ..@ assay.used : chr "RNA"
.. .. .. ..@ global : logi FALSE
.. .. .. ..@ stdev : num(0)
.. .. .. ..@ key : chr "spatial_"
.. .. .. ..@ jackstraw :Formal class 'JackStrawData' [package "SeuratObject"] with 4 slots
.. .. .. .. .. ..@ empirical.p.values : num[0 , 0 ]
.. .. .. .. .. ..@ fake.reduction.scores : num[0 , 0 ]
.. .. .. .. .. ..@ empirical.p.values.full: num[0 , 0 ]
.. .. .. .. .. ..@ overall.p.values : num[0 , 0 ]
.. .. .. ..@ misc : list()
..@ images : list()
..@ project.name: chr "SeuratProject"
..@ misc :List of 1
.. ..$ info:List of 2
.. .. ..$ demo :List of 4
.. .. .. ..$ img : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAAD6AAAAXACAAAAABnkOzKAAAgAElEQVR4AezB3ZJkSXKl12+rmp3jEVlVDQw4MxDw/Z9rmhShDE"| __truncated__
.. .. .. ..$ img_gg:List of 12
.. .. .. .. ..$ raster : 'raster' chr [1:1472, 1:4000] "#000000" "#000000" "#000000" "#000000" ...n .. .. .. .. ..$ x : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ y : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ width : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ height : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ just : chr "centre"
.. .. .. .. ..$ hjust : NULL
.. .. .. .. ..$ vjust : NULL
.. .. .. .. ..$ interpolate: logi FALSE
.. .. .. .. ..$ name : chr "GRID.rastergrob.768"
.. .. .. .. ..$ gp : list()
.. .. .. .. .. ..- attr(*, "class")= chr "gpar"
.. .. .. .. ..$ vp : NULL
.. .. .. .. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
.. .. .. ..$ size_x: int 55050
.. .. .. ..$ size_y: int 19906
.. .. ..$ WTH1092:List of 6
.. .. .. ..$ img : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAAD6AAAAWnCAAAAAByYdMoAAAgAElEQVR4AezBaa9l6WGe5/t537X22vOZ6lSdququrq5ukt2kRI"| __truncated__
.. .. .. ..$ img_gg :List of 12
.. .. .. .. ..$ raster : 'raster' chr [1:1447, 1:4000] "#000000" "#000000" "#000000" "#000000" ...n .. .. .. .. ..$ x : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ y : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ width : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ height : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ just : chr "centre"
.. .. .. .. ..$ hjust : NULL
.. .. .. .. ..$ vjust : NULL
.. .. .. .. ..$ interpolate: logi FALSE
.. .. .. .. ..$ name : chr "GRID.rastergrob.720"
.. .. .. .. ..$ gp : list()
.. .. .. .. .. ..- attr(*, "class")= chr "gpar"
.. .. .. .. ..$ vp : NULL
.. .. .. .. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
.. .. .. ..$ size_x : int 55128
.. .. .. ..$ size_y : int 19906
.. .. .. ..$ img_he : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAALuAAABD1CAIAAAB1YFaBAAAgAElEQVR4AezBwYHoxoJdybjJ8d8BreSnmEdA1SP7qzUGaIGIVT"| __truncated__
.. .. .. ..$ img_he_gg:List of 12
.. .. .. .. ..$ raster : 'raster' chr [1:4341, 1:12000] "#FFFFFF" "#FFFFFF" "#9B9BA1" "#7A7B83" ...n .. .. .. .. ..$ x : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ y : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ width : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ height : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ just : chr "centre"
.. .. .. .. ..$ hjust : NULL
.. .. .. .. ..$ vjust : NULL
.. .. .. .. ..$ interpolate: logi FALSE
.. .. .. .. ..$ name : chr "GRID.rastergrob.721"
.. .. .. .. ..$ gp : list()
.. .. .. .. .. ..- attr(*, "class")= chr "gpar"
.. .. .. .. ..$ vp : NULL
.. .. .. .. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
..@ version :Classes 'package_version', 'numeric_version' hidden list of 1
.. ..$ : int [1:3] 4 1 3
..@ commands : list()
..@ tools : list()
..@ assays :List of 1
.. ..$ RNA:Formal class 'Assay' [package "SeuratObject"] with 8 slots
.. .. .. ..@ counts :Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
.. .. .. .. .. ..@ i : int [1:47683687] 493 876 1115 1617 1838 1962 2169 2613 2959 4318 ...n .. .. .. .. .. ..@ p : int [1:59172] 0 117 197 271 402 469 520 669 754 890 ...n .. .. .. .. .. ..@ Dim : int [1:2] 32285 59171
.. .. .. .. .. ..@ Dimnames:List of 2
.. .. .. .. .. .. ..$ : chr [1:32285] "Xkr4" "Gm1992" "Gm19938" "Gm37381" ...n .. .. .. .. .. .. ..$ : chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n .. .. .. .. .. ..@ x : num [1:47683687] 1 1 1 1 1 1 1 1 1 1 ...n .. .. .. .. .. ..@ factors : list()
.. .. .. ..@ data :Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
.. .. .. .. .. ..@ i : int [1:47683687] 493 876 1115 1617 1838 1962 2169 2613 2959 4318 ...n .. .. .. .. .. ..@ p : int [1:59172] 0 117 197 271 402 469 520 669 754 890 ...n .. .. .. .. .. ..@ Dim : int [1:2] 32285 59171
.. .. .. .. .. ..@ Dimnames:List of 2
.. .. .. .. .. .. ..$ : chr [1:32285] "Xkr4" "Gm1992" "Gm19938" "Gm37381" ...n .. .. .. .. .. .. ..$ : chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n .. .. .. .. .. ..@ x : num [1:47683687] 3.93 3.93 3.93 3.93 3.93 ...n .. .. .. .. .. ..@ factors : list()
.. .. .. ..@ scale.data : num[0 , 0 ]
.. .. .. ..@ key : Named chr "rna_"
.. .. .. .. ..- attr(*, "names")= chr ""
.. .. .. ..@ assay.orig : NULL
.. .. .. ..@ var.features : logi(0)
.. .. .. ..@ meta.features:'data.frame': 32285 obs. of 0 variables
.. .. .. ..@ misc : list()
..@ meta.data :'data.frame': 59171 obs. of 12 variables:
.. ..$ orig.ident : chr [1:59171] "demo" "demo" "demo" "demo" ...n .. ..$ nCount_RNA : num [1:59171] 200 200 200 200 200 200 200 200 200 200 ...n .. ..$ nFeature_RNA : int [1:59171] 117 80 74 131 67 51 149 85 136 49 ...n .. ..$ percent.mito : num [1:59171] 0 0.5 1.5 0 0.5 0 0 0.5 0 0.5 ...n .. ..$ RNA_snn_res.0.2: chr [1:59171] "5" "6" "3" "1" ...n .. ..$ RNA_snn_res.0.5: chr [1:59171] "2" "3" "19" "12" ...n .. ..$ RNA_snn_res.0.8: chr [1:59171] "2" "16" "23" "15" ...n .. ..$ RNA_snn_res.1.1: chr [1:59171] "1" "7" "23" "17" ...n .. ..$ RNA_snn_res.1.4: chr [1:59171] "5" "2" "24" "17" ...n .. ..$ seurat_clusters: chr [1:59171] "5" "2" "24" "17" ...n .. ..$ Sample : chr [1:59171] "demo" "demo" "demo" "demo" ...n .. ..$ stim : chr [1:59171] "WTH1059" "WTH1059" "WTH1059" "WTH1059" ...n ..@ active.assay: chr "RNA"
..@ active.ident: Factor w/ 28 levels "0","1","10","11",..: 13 9 17 8 15 3 26 15 8 26 ...n .. ..- attr(*, "names")= chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n ..@ graphs : list()
..@ neighbors : list()
..@ reductions :List of 1
.. ..$ spatial:Formal class 'DimReduc' [package "SeuratObject"] with 9 slots
.. .. .. ..@ cell.embeddings : int [1:59171, 1:2] 34524 42576 44264 40223 36766 33959 26542 28047 39761 21306 ...n .. .. .. .. ..- attr(*, "dimnames")=List of 2
.. .. .. .. .. ..$ : chr [1:59171] "AAAGGGCGCTCCAGCCACGTGCCGATT" "ACCCAAACGAAGCGTAGAGACCGCCTT" "ACTTATCGCTAGACGGTAAGTTACTCG" "AGCGATTGCTGCTTTGATTCAGGTGGC" ...n .. .. .. .. .. ..$ : chr [1:2] "spatial_1" "spatial_2"
.. .. .. ..@ feature.loadings : num[0 , 0 ]
.. .. .. ..@ feature.loadings.projected: num[0 , 0 ]
.. .. .. ..@ assay.used : chr "RNA"
.. .. .. ..@ global : logi FALSE
.. .. .. ..@ stdev : num(0)
.. .. .. ..@ key : chr "spatial_"
.. .. .. ..@ jackstraw :Formal class 'JackStrawData' [package "SeuratObject"] with 4 slots
.. .. .. .. .. ..@ empirical.p.values : num[0 , 0 ]
.. .. .. .. .. ..@ fake.reduction.scores : num[0 , 0 ]
.. .. .. .. .. ..@ empirical.p.values.full: num[0 , 0 ]
.. .. .. .. .. ..@ overall.p.values : num[0 , 0 ]
.. .. .. ..@ misc : list()
..@ images : list()
..@ project.name: chr "SeuratProject"
..@ misc :List of 1
.. ..$ info:List of 2
.. .. ..$ demo :List of 4
.. .. .. ..$ img : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAAD6AAAAXACAAAAABnkOzKAAAgAElEQVR4AezB3ZJkSXKl12+rmp3jEVlVDQw4MxDw/Z9rmhShDE"| __truncated__
.. .. .. ..$ img_gg:List of 12
.. .. .. .. ..$ raster : 'raster' chr [1:1472, 1:4000] "#000000" "#000000" "#000000" "#000000" ...n .. .. .. .. ..$ x : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ y : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ width : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ height : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ just : chr "centre"
.. .. .. .. ..$ hjust : NULL
.. .. .. .. ..$ vjust : NULL
.. .. .. .. ..$ interpolate: logi FALSE
.. .. .. .. ..$ name : chr "GRID.rastergrob.768"
.. .. .. .. ..$ gp : list()
.. .. .. .. .. ..- attr(*, "class")= chr "gpar"
.. .. .. .. ..$ vp : NULL
.. .. .. .. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
.. .. .. ..$ size_x: int 55050
.. .. .. ..$ size_y: int 19906
.. .. ..$ WTH1092:List of 6
.. .. .. ..$ img : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAAD6AAAAWnCAAAAAByYdMoAAAgAElEQVR4AezBaa9l6WGe5/t537X22vOZ6lSdququrq5ukt2kRI"| __truncated__
.. .. .. ..$ img_gg :List of 12
.. .. .. .. ..$ raster : 'raster' chr [1:1447, 1:4000] "#000000" "#000000" "#000000" "#000000" ...n .. .. .. .. ..$ x : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ y : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ width : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ height : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ just : chr "centre"
.. .. .. .. ..$ hjust : NULL
.. .. .. .. ..$ vjust : NULL
.. .. .. .. ..$ interpolate: logi FALSE
.. .. .. .. ..$ name : chr "GRID.rastergrob.720"
.. .. .. .. ..$ gp : list()
.. .. .. .. .. ..- attr(*, "class")= chr "gpar"
.. .. .. .. ..$ vp : NULL
.. .. .. .. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
.. .. .. ..$ size_x : int 55128
.. .. .. ..$ size_y : int 19906
.. .. .. ..$ img_he : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAALuAAABD1CAIAAAB1YFaBAAAgAElEQVR4AezBwYHoxoJdybjJ8d8BreSnmEdA1SP7qzUGaIGIVT"| __truncated__
.. .. .. ..$ img_he_gg:List of 12
.. .. .. .. ..$ raster : 'raster' chr [1:4341, 1:12000] "#FFFFFF" "#FFFFFF" "#9B9BA1" "#7A7B83" ...n .. .. .. .. ..$ x : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ y : 'simpleUnit' num 0.5npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ width : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ height : 'simpleUnit' num 1npc
.. .. .. .. .. ..- attr(*, "unit")= int 0
.. .. .. .. ..$ just : chr "centre"
.. .. .. .. ..$ hjust : NULL
.. .. .. .. ..$ vjust : NULL
.. .. .. .. ..$ interpolate: logi FALSE
.. .. .. .. ..$ name : chr "GRID.rastergrob.721"
.. .. .. .. ..$ gp : list()
.. .. .. .. .. ..- attr(*, "class")= chr "gpar"
.. .. .. .. ..$ vp : NULL
.. .. .. .. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
..@ version :Classes 'package_version', 'numeric_version' hidden list of 1
.. ..$ : int [1:3] 4 1 3
..@ commands : list()
..@ tools : list()
样本正常 merge 之后就可以选择正常的 harmony 或 cca 进行去批次和降维聚类分析
R
obj.integrated <- NormalizeData(obj.integrated, normalization.method = "LogNormalize", scale.factor = 10000)
obj.integrated <- FindVariableFeatures(obj.integrated, selection.method = "vst", nfeatures = 2000)
obj.integrated <- ScaleData(obj.integrated, verbose = FALSE)
obj.integrated <- RunPCA(obj.integrated, npcs = 20, verbose = FALSE)R
#harmony去批次分析
obj.integrated <- obj.integrated %>% RunHarmony("stim", plot_convergence = TRUE)
#cca去批次分析
#obj.anchors <- FindIntegrationAnchors(object.list = ob.list, dims = 1:dims)
#obj.integrated <- IntegrateData(anchorset = obj.anchors, dims = 1:dims)output
Warning message:
“Quick-TRANSfer stage steps exceeded maximum (= 2958550)”
Harmony 1/10
Harmony 2/10
Harmony 3/10
Harmony 4/10
Harmony 5/10
Harmony 6/10
Harmony 7/10
Harmony 8/10
Harmony 9/10
Harmony 10/10
Warning message:
“Invalid name supplied, making object name syntactically valid. New object name is Seurat..ProjectDim.RNA.harmony; see ?make.names for more details on syntax validity”
“Quick-TRANSfer stage steps exceeded maximum (= 2958550)”
Harmony 1/10
Harmony 2/10
Harmony 3/10
Harmony 4/10
Harmony 5/10
Harmony 6/10
Harmony 7/10
Harmony 8/10
Harmony 9/10
Harmony 10/10
Warning message:
“Invalid name supplied, making object name syntactically valid. New object name is Seurat..ProjectDim.RNA.harmony; see ?make.names for more details on syntax validity”
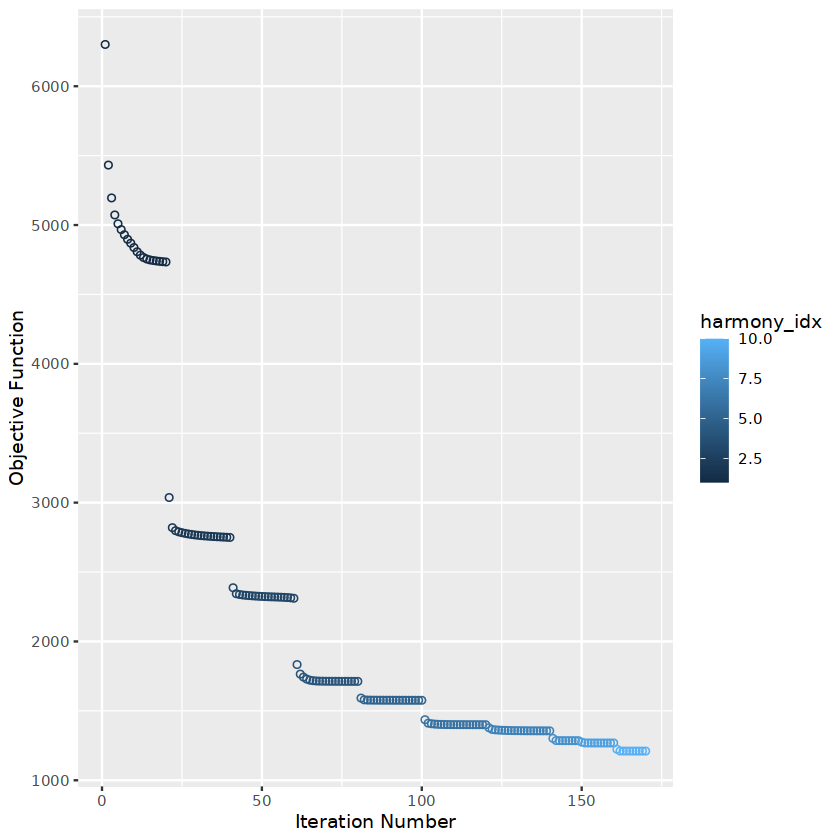
R
obj.integrated <- RunUMAP(obj.integrated, reduction = "harmony", dims = 1:20)
obj.integrated <- RunTSNE(obj.integrated, reduction = "harmony", dims = 1:20)
obj.integrated <- FindNeighbors(obj.integrated,reduction = "harmony", dims = 1:20)
obj.integrated <- FindClusters(obj.integrated, resolution = 0.8)output
Warning message:
“The default method for RunUMAP has changed from calling Python UMAP via reticulate to the R-native UWOT using the cosine metric
To use Python UMAP via reticulate, set umap.method to 'umap-learn' and metric to 'correlation'
This message will be shown once per session”
12:36:57 UMAP embedding parameters a = 0.9922 b = 1.112
12:36:57 Read 59171 rows and found 20 numeric columns
12:36:57 Using Annoy for neighbor search, n_neighbors = 30
12:36:57 Building Annoy index with metric = cosine, n_trees = 50
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
|
12:37:05 Writing NN index file to temp file /tmp/RtmpE8Pqe6/file3eca9fadcb
12:37:05 Searching Annoy index using 1 thread, search_k = 3000
12:37:29 Annoy recall = 100%
12:37:29 Commencing smooth kNN distance calibration using 1 thread
with target n_neighbors = 30
12:37:33 Initializing from normalized Laplacian + noise (using irlba)
12:37:41 Commencing optimization for 200 epochs, with 2742632 positive edges
12:38:24 Optimization finished
Computing nearest neighbor graph
Computing SNN
Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 59171
Number of edges: 2182636
Running Louvain algorithm...n Maximum modularity in 10 random starts: 0.9357
Number of communities: 31
Elapsed time: 17 seconds
“The default method for RunUMAP has changed from calling Python UMAP via reticulate to the R-native UWOT using the cosine metric
To use Python UMAP via reticulate, set umap.method to 'umap-learn' and metric to 'correlation'
This message will be shown once per session”
12:36:57 UMAP embedding parameters a = 0.9922 b = 1.112
12:36:57 Read 59171 rows and found 20 numeric columns
12:36:57 Using Annoy for neighbor search, n_neighbors = 30
12:36:57 Building Annoy index with metric = cosine, n_trees = 50
0% 10 20 30 40 50 60 70 80 90 100%
[----|----|----|----|----|----|----|----|----|----|
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
*
|
12:37:05 Writing NN index file to temp file /tmp/RtmpE8Pqe6/file3eca9fadcb
12:37:05 Searching Annoy index using 1 thread, search_k = 3000
12:37:29 Annoy recall = 100%
12:37:29 Commencing smooth kNN distance calibration using 1 thread
with target n_neighbors = 30
12:37:33 Initializing from normalized Laplacian + noise (using irlba)
12:37:41 Commencing optimization for 200 epochs, with 2742632 positive edges
12:38:24 Optimization finished
Computing nearest neighbor graph
Computing SNN
Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
Number of nodes: 59171
Number of edges: 2182636
Running Louvain algorithm...n Maximum modularity in 10 random starts: 0.9357
Number of communities: 31
Elapsed time: 17 seconds
R
head(obj.integrated)| orig.ident | nCount_RNA | nFeature_RNA | percent.mito | RNA_snn_res.0.2 | RNA_snn_res.0.5 | RNA_snn_res.0.8 | RNA_snn_res.1.1 | RNA_snn_res.1.4 | seurat_clusters | Sample | stim | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| <chr> | <dbl> | <int> | <dbl> | <chr> | <chr> | <fct> | <chr> | <chr> | <fct> | <chr> | <chr> | |
| AAAGGGCGCTCCAGCCACGTGCCGATT | demo | 200 | 117 | 0.0 | 5 | 2 | 1 | 1 | 5 | 1 | demo | WTH1059 |
| ACCCAAACGAAGCGTAGAGACCGCCTT | demo | 200 | 80 | 0.5 | 6 | 3 | 11 | 7 | 2 | 11 | demo | WTH1059 |
| ACTTATCGCTAGACGGTAAGTTACTCG | demo | 200 | 74 | 1.5 | 3 | 19 | 19 | 23 | 24 | 19 | demo | WTH1059 |
| AGCGATTGCTGCTTTGATTCAGGTGGC | demo | 200 | 131 | 0.0 | 1 | 12 | 19 | 17 | 17 | 19 | demo | WTH1059 |
| AGCGCTGTACACAGCTGAGACCGCCTT | demo | 200 | 67 | 0.5 | 11 | 16 | 24 | 22 | 22 | 24 | demo | WTH1059 |
| ATCGCCTTACTGGCCAGCATGGCGTAC | demo | 200 | 51 | 0.0 | 4 | 5 | 4 | 12 | 16 | 4 | demo | WTH1059 |
| ATTACCTCGAGCTCGGTAAGTTACTCG | demo | 200 | 149 | 0.0 | 3 | 0 | 0 | 2 | 6 | 0 | demo | WTH1059 |
| CAACGGCATGCCTAGGAAGACCGCCTT | demo | 200 | 85 | 0.5 | 11 | 16 | 24 | 22 | 22 | 24 | demo | WTH1059 |
| CCACTTGGCTTTGATCGCGTGCCGATT | demo | 200 | 136 | 0.0 | 1 | 12 | 19 | 17 | 17 | 19 | demo | WTH1059 |
| CCTCAGTGCTCTCAAGTTTCAGGTGGC | demo | 200 | 49 | 0.5 | 3 | 0 | 19 | 2 | 6 | 19 | demo | WTH1059 |
结果可视化
多样本的可视化可以参考 seekspace_of_Seurat.ipynb
R
options(repr.plot.height=7, repr.plot.width=14)
DimPlot(obj.integrated, reduction = 'umap', group.by=c("stim","RNA_snn_res.0.8"))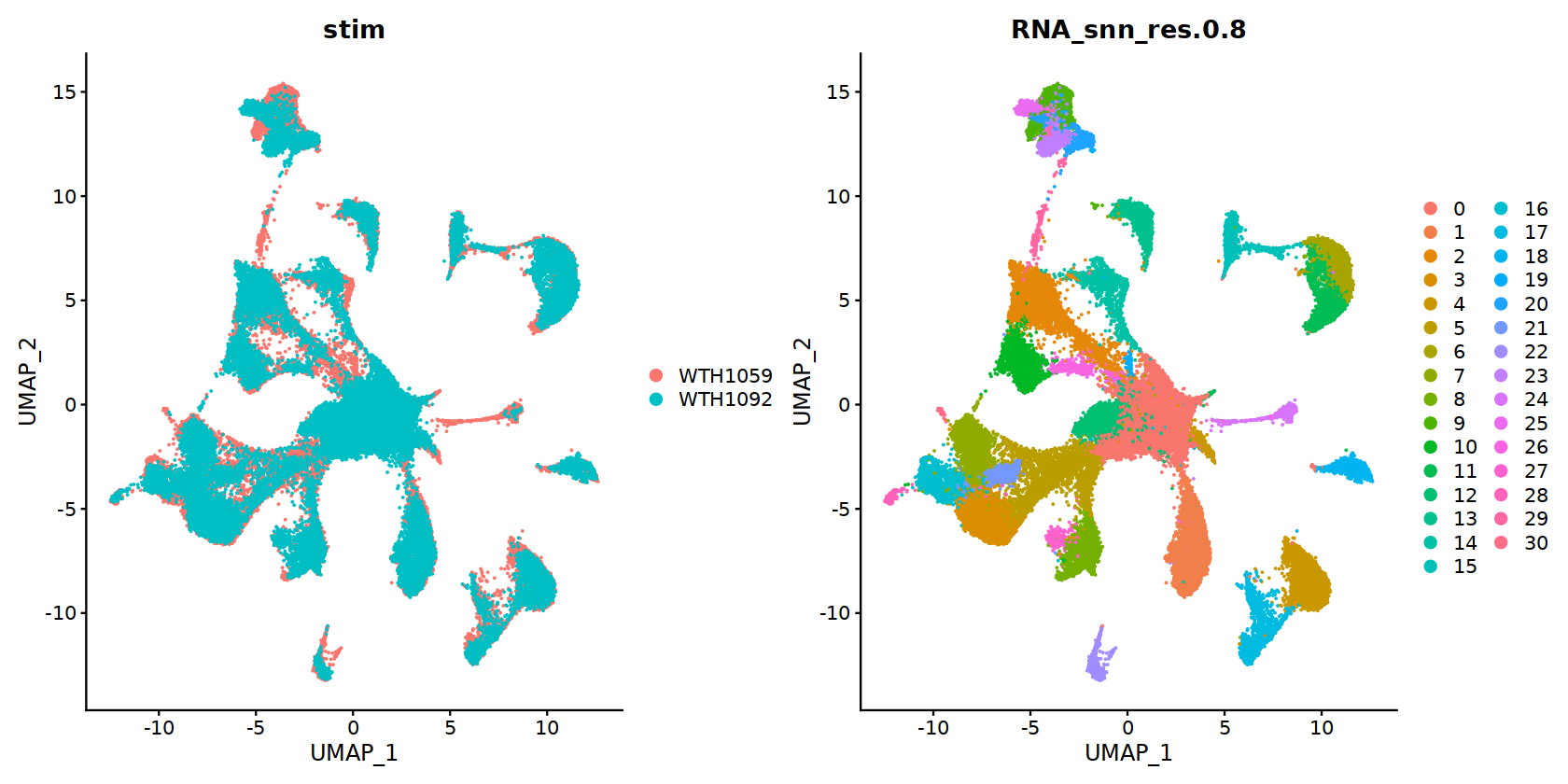
当我们把 DimPlot 函数中的 reduction 参数设置为 "spatial" 的时候,我们即可将细胞绘制在空间水平上。
R
options(repr.plot.height=7, repr.plot.width=15)
DimPlot(subset(obj.integrated,subset=stim=="WTH1092"), reduction = 'spatial',pt.size = 1)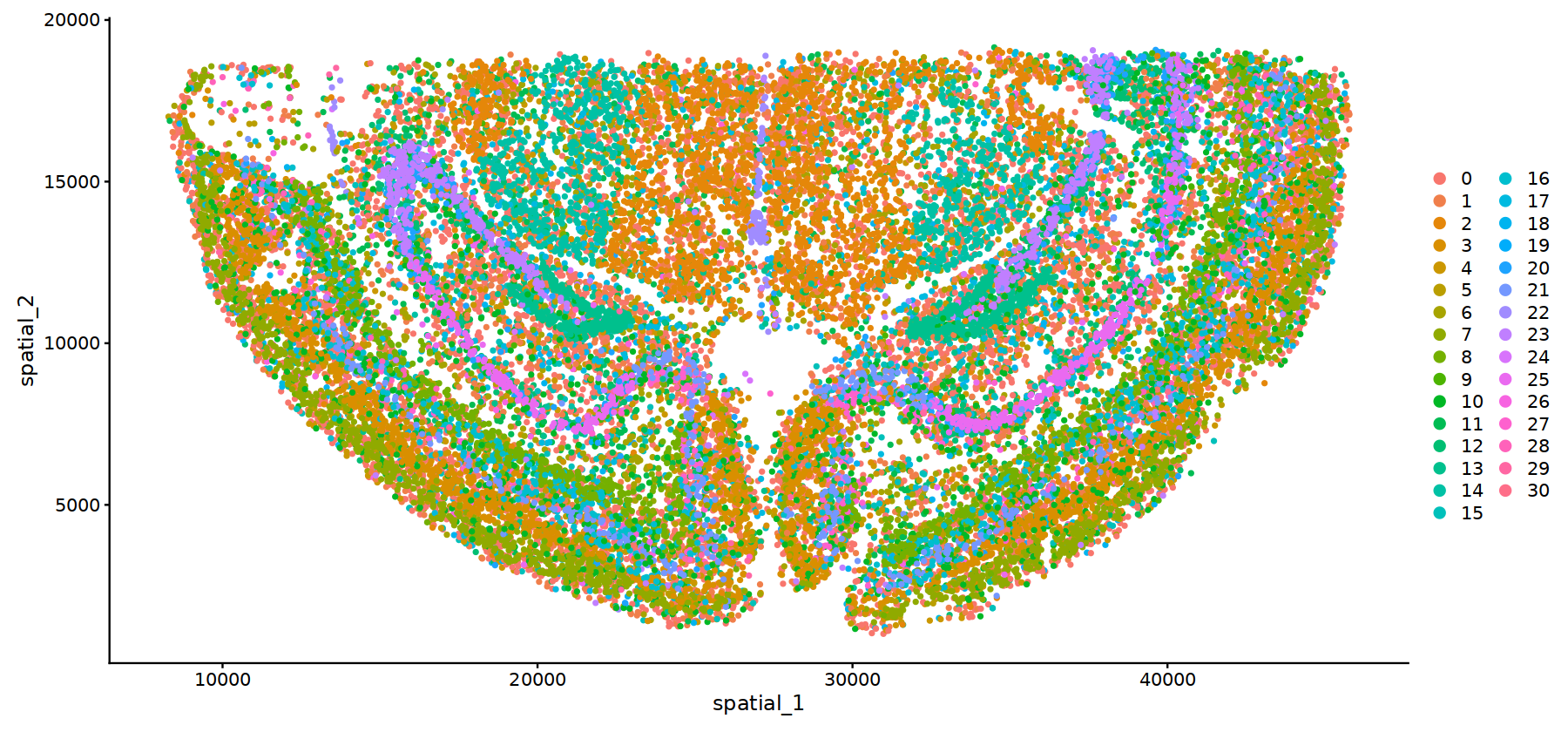
R
options(repr.plot.height=7, repr.plot.width=15)
DimPlot(subset(obj.integrated,subset=stim=="WTH1059"), reduction = 'spatial',pt.size = 1)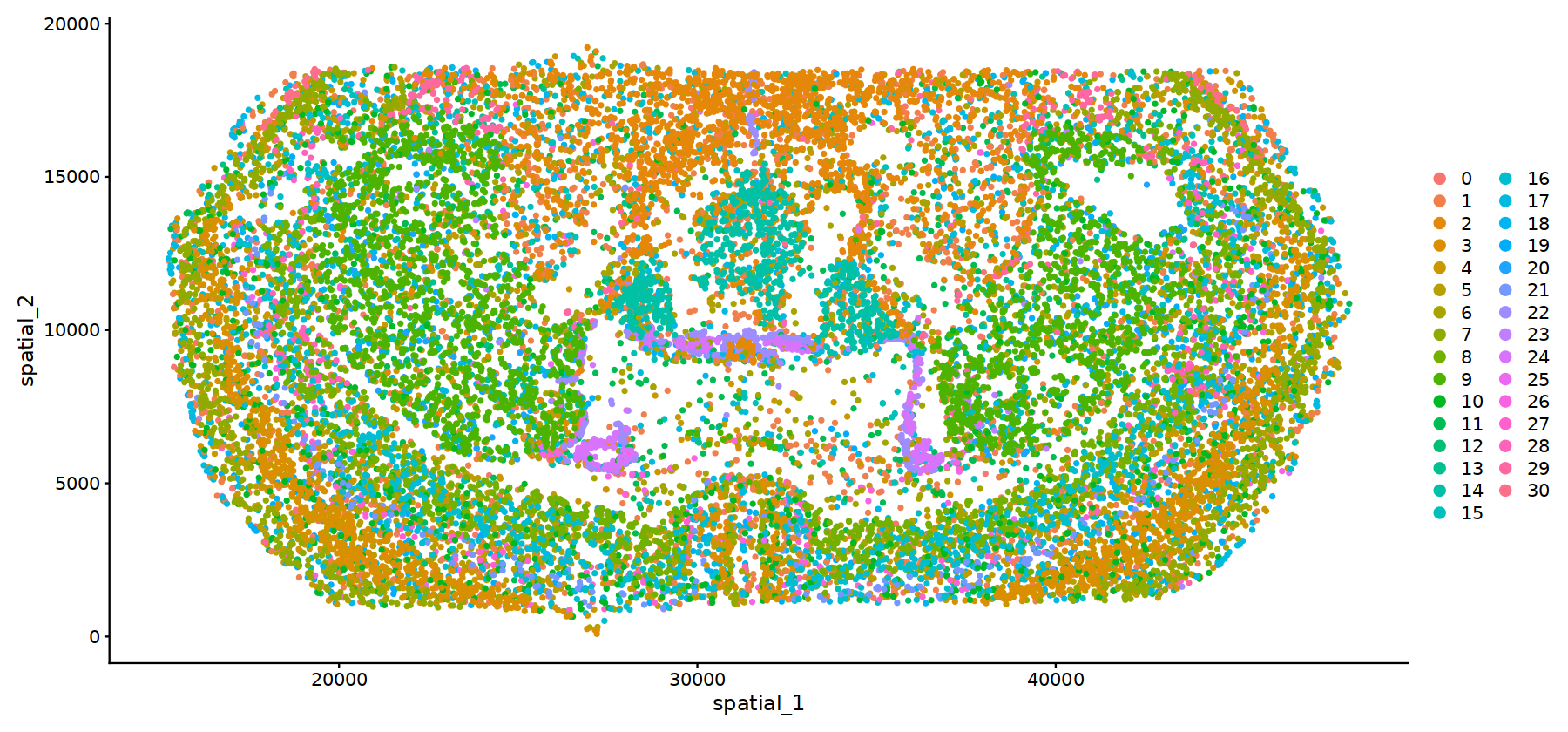
R
#自定义了ImageSpacePlot和FeatureSpacePlot两个函数绘制细胞在空间上的分群信息和细胞连续变量的指标R
######################绘图细胞分群函数
ImageSpacePlot = function(obj, group_by, type="DAPI", sample=names(obj@misc$info)[1], size=1, alpha=1,color=MYCOLOR){
MYCOLOR=c(
"#6394ce", "#2a4c87", "#eed500", "#ed5858",
"#f6cbc2", "#f5a2a2", "#3ca676", "#6cc9d8",
"#ef4db0", "#992269", "#bcb34a", "#74acf3",
"#3e275b", "#fbec7e", "#ec4d3d", "#ee807e",
"#f7bdb5", "#dbdde6", "#f591e1", "#51678c",
"#2fbcd3", "#80cfc3", "#fbefd1", "#edb8b5",
"#5678a8", "#2fb290", "#a6b5cd", "#90d1c1",
"#a4e0ea", "#837fd3", "#5dce8b", "#c5cdd9",
"#f9e2d6", "#c64ea4", "#b2dfd6", "#dbdfe7",
"#dff2ec", "#cce8f3", "#e74d51", "#f7c9c4",
"#f29c81", "#c9e6e0", "#c1c5de", "#750000"
)
raster_type <- switch(type,
HE = "img_he_gg",
DAPI = "img_gg",
stop("Invalid type. Must be 'HE' or 'DAPI'.")
)
spatial_coord1 <- as.data.frame(obj[[group_by]])
colnames(spatial_coord1) <- group_by
spatial_coord2 <- as.data.frame(obj@reductions$spatial@cell.embeddings)
spatial_coord <-cbind(spatial_coord2,spatial_coord1)
ImageSpacePlot <- ggplot2::ggplot() + ggplot2::annotation_custom(grob = obj@misc$info[[sample]][[raster_type]],
xmin = 0, xmax = obj@misc$info[[sample]]$size_x,
ymin = 0, ymax = obj@misc$info[[sample]]$size_y) +
ggplot2::geom_point(data = spatial_coord, ggplot2::aes(x = spatial_1,y = spatial_2, color = !!sym(group_by),
fill = !!sym(group_by)), size=size, alpha=alpha)+
labs(size = group_by) + guides(alpha = "none")+
ggplot2::theme_classic()+
scale_color_manual(values = color)+ coord_fixed()
return(ImageSpacePlot)
}
###################绘图基因表达函数
FeatureSpacePlot = function(obj, feature, type="DAPI", sample=names(obj@misc$info)[1], size=1, alpha=c(1,1),color=c("lightgrey","blue")){
raster_type <- switch(type,
HE = "img_he_gg",
DAPI = "img_gg",
stop("Invalid type. Must be 'HE' or 'DAPI'.")
)
spatial_coord1 <- as.data.frame(obj@reductions$spatial@cell.embeddings)
spatial_coord2 <- FetchData(obj,feature)
colnames(spatial_coord2) <- feature
spatial_coord <-cbind(spatial_coord1,spatial_coord2)
FeatureSpacePlot <-ggplot2::ggplot() + ggplot2::annotation_custom(grob = obj@misc$info[[sample]][[raster_type]],
xmin = 0, xmax = obj@misc$info[[sample]]$size_x, ymin = 0, ymax = obj@misc$info[[sample]]$size_y) +
ggplot2::geom_point(data = spatial_coord, ggplot2::aes(x = spatial_1, y = spatial_2,color = !!sym(feature),alpha = !!sym(feature)), size=size)+
labs(color = feature)+
guides(alpha = "none")+
ggplot2::theme_classic()+
ggplot2::scale_alpha_continuous(range=alpha)+
scale_color_gradient(low=color[1],high = color[2])+ coord_fixed()
return(FeatureSpacePlot)
}R
# DAPI背景的细胞分群图
options(repr.plot.height=7, repr.plot.width=15)
ImageSpacePlot(subset(obj.integrated,subset=stim=="WTH1092"), group_by = "RNA_snn_res.0.8",type="DAPI",size=0.7,sample="WTH1092")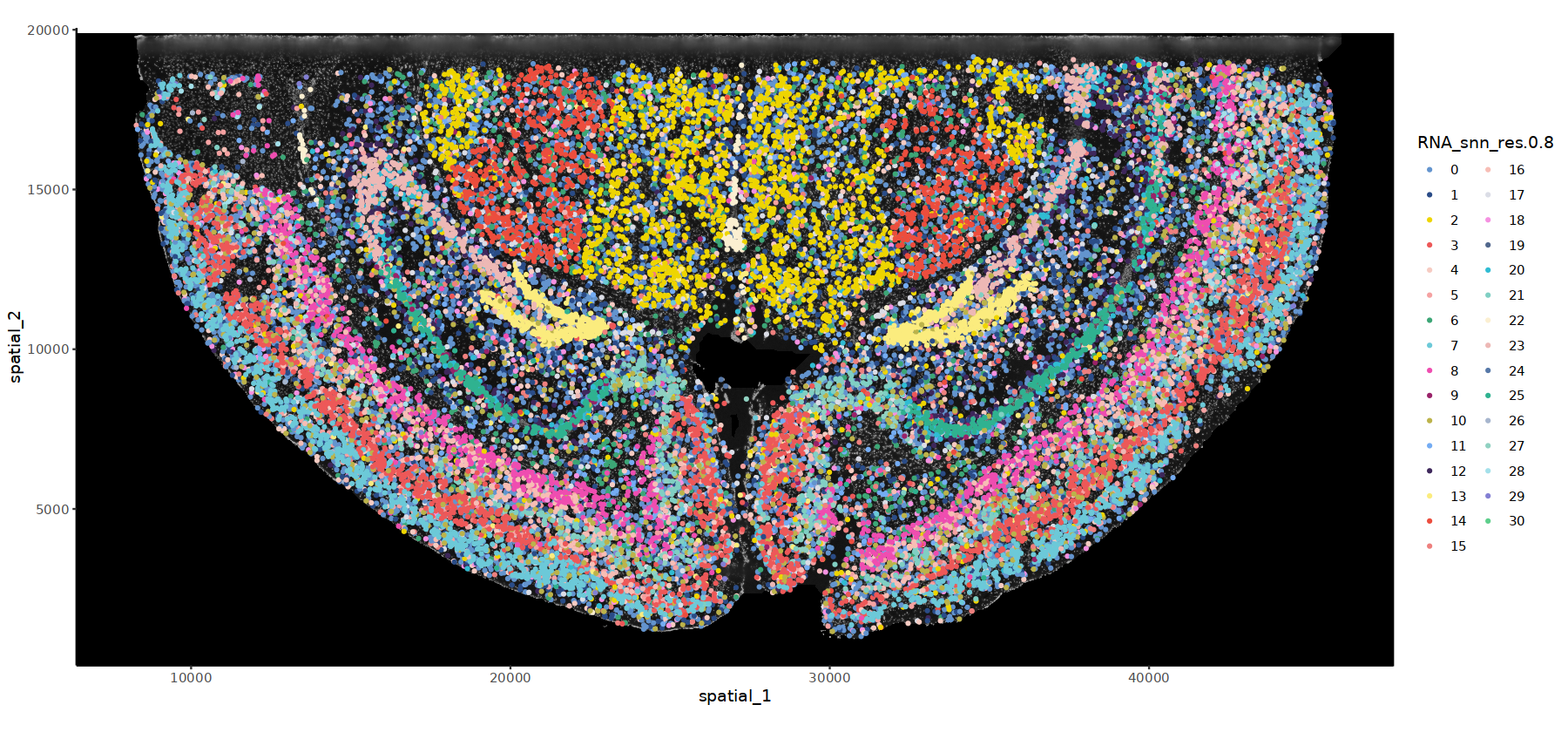
R
# DAPI背景的细胞分群图
options(repr.plot.height=7, repr.plot.width=15)
ImageSpacePlot(subset(obj.integrated,subset=stim=="WTH1059"), group_by = "RNA_snn_res.0.8",type="DAPI",size=0.7,sample="WTH1059")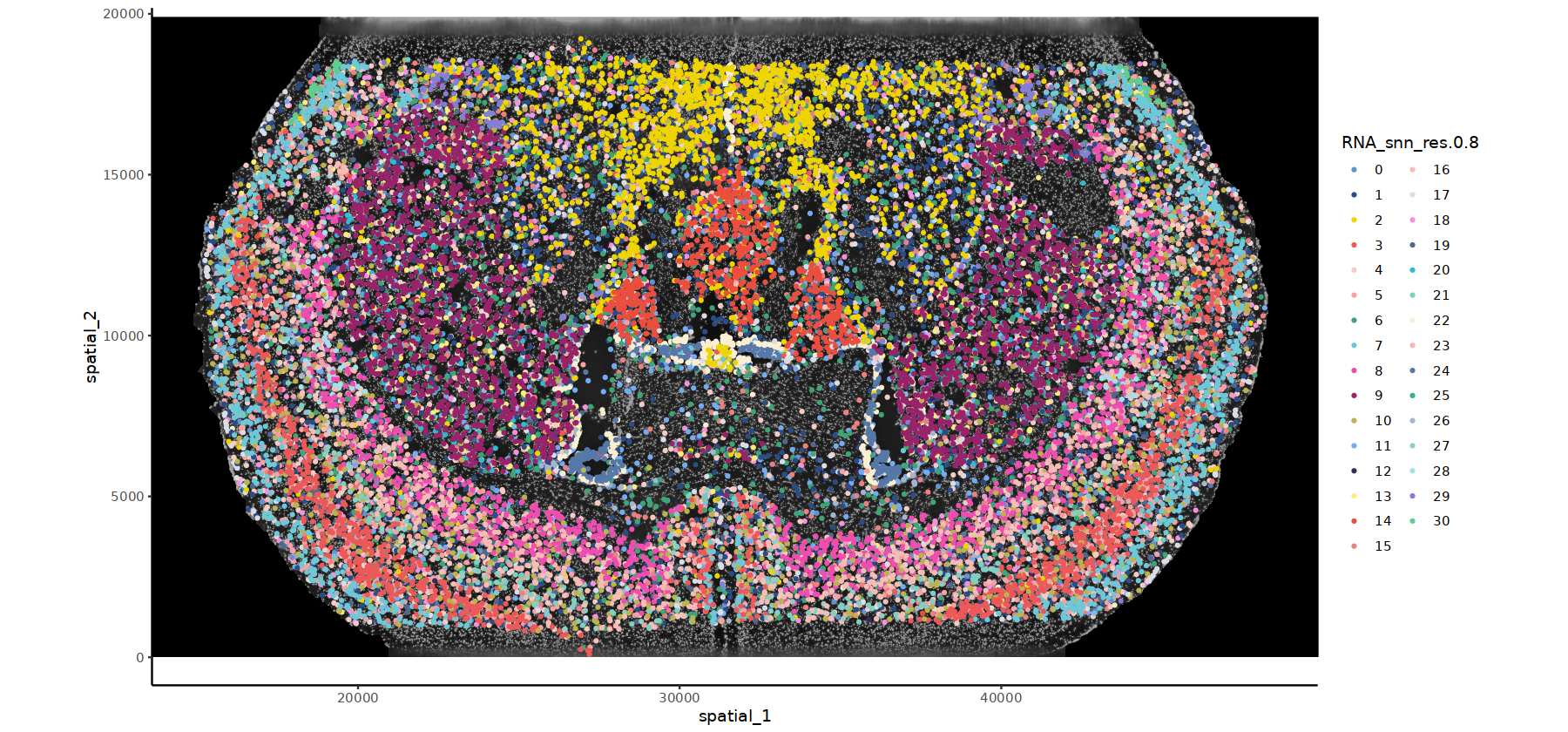
R
