SeekSpace 空间转录组可视化:基础绘图与参数配置
时长: 10 分钟
字数: 1.8k 字
更新: 2026-02-28
阅读: 0 次
R
# 加载R包
suppressMessages({
library(Seurat)
library(tidyverse)
library(ggplot2)
})R
# 读取数据
input <- readRDS("/home/demo-seekgene-com/workspace/data/AY1756127460073/input.rds")
meta <- read.table("/home/demo-seekgene-com/workspace/data/AY1756127460073/meta.tsv", header=TRUE, sep="\t", row.names = 1)
obj <- AddMetaData(input,meta)
head(obj@meta.data)
str(obj@misc$info)| orig.ident | nCount_RNA | nFeature_RNA | mito | Sample | raw_Sample | resolution.0.8_d30 | resolution.0.5_d30 | CellAnnotation | all | Sub_CellType | Main_CellType | SS | spatial_clustering | auto | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| <chr> | <int> | <int> | <dbl> | <chr> | <chr> | <int> | <int> | <chr> | <chr> | <chr> | <chr> | <chr> | <chr> | <chr> | |
| AAGGAATGCTGATTCGTTTCTGCGCTC | XYRD_WTH1092_expression | 300 | 163 | 4.6667 | WTH1092_expression | XYRD_WTH1092_expression | 0 | 0 | Neurons | all | Unknown | Unknown | undefined | undefined | Neurons |
| ACCGTTCTACAGCCATGTAGTCATACG | XYRD_WTH1092_expression | 300 | 204 | 1.0000 | WTH1092_expression | XYRD_WTH1092_expression | 6 | 6 | Astrocytes | all | Endo | Endo | undefined | undefined | Astrocytes |
| ACTCTCGTACGTCGTCCAAGGCACTAT | XYRD_WTH1092_expression | 300 | 170 | 1.6667 | WTH1092_expression | XYRD_WTH1092_expression | 0 | 0 | Neurons | all | Unknown | Unknown | undefined | undefined | Neurons |
| AGCTACAATGGAAAGGTGGACTGTGGA | XYRD_WTH1092_expression | 300 | 179 | 3.3333 | WTH1092_expression | XYRD_WTH1092_expression | 19 | 19 | Neurons | all | Ext_Thal_Unk | Ext | undefined | undefined | Neurons |
| AGGACGACGAGATAAACTAGTCATACG | XYRD_WTH1092_expression | 300 | 166 | 4.6667 | WTH1092_expression | XYRD_WTH1092_expression | 19 | 19 | Neurons | all | Ext_Thal_Unk | Ext | undefined | undefined | Neurons |
| AGGCATTTACCGAACTGTTCTGCGCTC | XYRD_WTH1092_expression | 300 | 231 | 0.3333 | WTH1092_expression | XYRD_WTH1092_expression | 2 | 1 | Neurons | all | Inh | Inh | undefined | undefined | Neurons |
List of 1
$ WTH1092_expression:List of 6
..$ img : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAAD6AAAAWnCAAAAAByYdMoAAAgAElEQVR4AezBaa9l6WGe5/t537X22vOZ6lSdququrh5IdpMiRU"| __truncated__
..$ img_gg :List of 12
.. ..$ raster : 'raster' chr [1:1447, 1:4000] "#000000" "#000000" "#000000" "#000000" ...
.. ..$ x : 'simpleUnit' num 0.5npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ y : 'simpleUnit' num 0.5npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ width : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ height : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ just : chr "centre"
.. ..$ hjust : NULL
.. ..$ vjust : NULL
.. ..$ interpolate: logi FALSE
.. ..$ name : chr "GRID.rastergrob.708"
.. ..$ gp : list()
.. .. ..- attr(*, "class")= chr "gpar"
.. ..$ vp : NULL
.. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
..$ size_x : int 55128
..$ size_y : int 19906
..$ img_he : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAALuAAABD1CAIAAAB1YFaBAAAgAElEQVR4AezBwYHoxoJdybjJ8d8BreSnmEdA1SP7qzUGaIGIVT"| __truncated__
..$ img_he_gg:List of 12
.. ..$ raster : 'raster' chr [1:4341, 1:12000] "#FFFFFF" "#FFFFFF" "#9B9BA1" "#7A7B83" ...
.. ..$ x : 'simpleUnit' num 0.5npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ y : 'simpleUnit' num 0.5npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ width : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ height : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ just : chr "centre"
.. ..$ hjust : NULL
.. ..$ vjust : NULL
.. ..$ interpolate: logi FALSE
.. ..$ name : chr "GRID.rastergrob.709"
.. ..$ gp : list()
.. .. ..- attr(*, "class")= chr "gpar"
.. ..$ vp : NULL
.. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
R
######################base64转png图函数
Base64ToPng <- function(obj, width_px = 1000) {
for (i in seq_along(obj@misc$info)) {
for (j in c("img", "img_he")) {
val <- obj@misc$info[[i]][[j]]
if (is.null(val) || is.na(val)) next
b64 <- stringr::str_remove(val, '^data:[^;]*;base64,')
img_raw <- base64enc::base64decode(b64)
grob <- magick::image_read(img_raw) %>%
magick::image_resize(sprintf("%d>", width_px)) %>%
grid::rasterGrob(x = 0, y = 0,
width = grid::unit(1, "npc"), height = grid::unit(1, "npc"),
just = c(0, 0), interpolate = TRUE)
key <- if (j == "img") "img_gg" else "img_he_gg"
obj@misc$info[[i]][[key]] <<- grob
}
}
invisible(obj)
}
######################绘图细胞分群函数
ImageSpacePlot = function(obj, group_by, type="DAPI", sample=names(obj@misc$info)[1], size=1, alpha=1,color=MYCOLOR){
MYCOLOR=c(
"#6394ce", "#2a4c87", "#eed500", "#ed5858",
"#f6cbc2", "#f5a2a2", "#3ca676", "#6cc9d8",
"#ef4db0", "#992269", "#bcb34a", "#74acf3",
"#3e275b", "#fbec7e", "#ec4d3d", "#ee807e",
"#f7bdb5", "#dbdde6", "#f591e1", "#51678c",
"#2fbcd3", "#80cfc3", "#fbefd1", "#edb8b5",
"#5678a8", "#2fb290", "#a6b5cd", "#90d1c1",
"#a4e0ea", "#837fd3", "#5dce8b", "#c5cdd9",
"#f9e2d6", "#c64ea4", "#b2dfd6", "#dbdfe7",
"#dff2ec", "#cce8f3", "#e74d51", "#f7c9c4",
"#f29c81", "#c9e6e0", "#c1c5de", "#750000"
)
raster_type <- switch(type,
HE = "img_he_gg",
DAPI = "img_gg",
stop("Invalid type. Must be 'HE' or 'DAPI'.")
)
spatial_coord1 <- as.data.frame(obj[[group_by]])
colnames(spatial_coord1) <- group_by
spatial_coord2 <- as.data.frame(obj@reductions$spatial@cell.embeddings)
spatial_coord <-cbind(spatial_coord2,spatial_coord1)
ImageSpacePlot <- ggplot2::ggplot() + ggplot2::annotation_custom(grob = obj@misc$info[[sample]][[raster_type]],
xmin = 0, xmax = obj@misc$info[[sample]]$size_x,
ymin = 0, ymax = obj@misc$info[[sample]]$size_y) +
ggplot2::geom_point(data = spatial_coord, ggplot2::aes(x = spatial_1,y = spatial_2, color = !!sym(group_by),
fill = !!sym(group_by)), size=size, alpha=alpha)+
labs(size = group_by) + guides(alpha = "none")+
ggplot2::theme_classic()+
scale_color_manual(values = color)+ coord_fixed()
return(ImageSpacePlot)
}
###################绘图基因表达函数
FeatureSpacePlot = function(obj, feature, type="DAPI", sample=names(obj@misc$info)[1], size=1, alpha=c(1,1),color=c("lightgrey","blue")){
raster_type <- switch(type,
HE = "img_he_gg",
DAPI = "img_gg",
stop("Invalid type. Must be 'HE' or 'DAPI'.")
)
spatial_coord1 <- as.data.frame(obj@reductions$spatial@cell.embeddings)
spatial_coord2 <- FetchData(obj,feature)
colnames(spatial_coord2) <- feature
spatial_coord <-cbind(spatial_coord1,spatial_coord2)
FeatureSpacePlot <-ggplot2::ggplot() + ggplot2::annotation_custom(grob = obj@misc$info[[sample]][[raster_type]],
xmin = 0, xmax = obj@misc$info[[sample]]$size_x, ymin = 0, ymax = obj@misc$info[[sample]]$size_y) +
ggplot2::geom_point(data = spatial_coord, ggplot2::aes(x = spatial_1, y = spatial_2,color = !!sym(feature),alpha = !!sym(feature)), size=size)+
labs(color = feature)+
guides(alpha = "none")+
ggplot2::theme_classic()+
ggplot2::scale_alpha_continuous(range=alpha)+
scale_color_gradient(low=color[1],high = color[2])+ coord_fixed()
return(FeatureSpacePlot)
}R
# 将base64转成PNG图
Base64ToPng(obj)
str(obj@misc$info) # 增加了img_gg(DAPI png图)和img_he_gg(HE png图)output
List of 1
$ WTH1092_expression:List of 6
..$ img : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAAD6AAAAWnCAAAAAByYdMoAAAgAElEQVR4AezBaa9l6WGe5/t537X22vOZ6lSdququrh5IdpMiRU"| __truncated__
..$ img_gg :List of 12
.. ..$ raster : 'raster' chr [1:362, 1:1000] "#000000ff" "#000000ff" "#000000ff" "#000000ff" ...n .. ..$ x : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ y : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ width : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ height : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ just : num [1:2] 0 0
.. ..$ hjust : NULL
.. ..$ vjust : NULL
.. ..$ interpolate: logi TRUE
.. ..$ name : chr "GRID.rastergrob.11"
.. ..$ gp : list()
.. .. ..- attr(*, "class")= chr "gpar"
.. ..$ vp : NULL
.. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
..$ size_x : int 55128
..$ size_y : int 19906
..$ img_he : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAALuAAABD1CAIAAAB1YFaBAAAgAElEQVR4AezBwYHoxoJdybjJ8d8BreSnmEdA1SP7qzUGaIGIVT"| __truncated__
..$ img_he_gg:List of 12
.. ..$ raster : 'raster' chr [1:362, 1:1000] "#888890ff" "#63636bff" "#575659ff" "#393832ff" ...n .. ..$ x : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ y : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ width : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ height : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ just : num [1:2] 0 0
.. ..$ hjust : NULL
.. ..$ vjust : NULL
.. ..$ interpolate: logi TRUE
.. ..$ name : chr "GRID.rastergrob.12"
.. ..$ gp : list()
.. .. ..- attr(*, "class")= chr "gpar"
.. ..$ vp : NULL
.. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
$ WTH1092_expression:List of 6
..$ img : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAAD6AAAAWnCAAAAAByYdMoAAAgAElEQVR4AezBaa9l6WGe5/t537X22vOZ6lSdququrh5IdpMiRU"| __truncated__
..$ img_gg :List of 12
.. ..$ raster : 'raster' chr [1:362, 1:1000] "#000000ff" "#000000ff" "#000000ff" "#000000ff" ...n .. ..$ x : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ y : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ width : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ height : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ just : num [1:2] 0 0
.. ..$ hjust : NULL
.. ..$ vjust : NULL
.. ..$ interpolate: logi TRUE
.. ..$ name : chr "GRID.rastergrob.11"
.. ..$ gp : list()
.. .. ..- attr(*, "class")= chr "gpar"
.. ..$ vp : NULL
.. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
..$ size_x : int 55128
..$ size_y : int 19906
..$ img_he : chr "data:;base64,iVBORw0KGgoAAAANSUhEUgAALuAAABD1CAIAAAB1YFaBAAAgAElEQVR4AezBwYHoxoJdybjJ8d8BreSnmEdA1SP7qzUGaIGIVT"| __truncated__
..$ img_he_gg:List of 12
.. ..$ raster : 'raster' chr [1:362, 1:1000] "#888890ff" "#63636bff" "#575659ff" "#393832ff" ...n .. ..$ x : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ y : 'simpleUnit' num 0npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ width : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ height : 'simpleUnit' num 1npc
.. .. ..- attr(*, "unit")= int 0
.. ..$ just : num [1:2] 0 0
.. ..$ hjust : NULL
.. ..$ vjust : NULL
.. ..$ interpolate: logi TRUE
.. ..$ name : chr "GRID.rastergrob.12"
.. ..$ gp : list()
.. .. ..- attr(*, "class")= chr "gpar"
.. ..$ vp : NULL
.. ..- attr(*, "class")= chr [1:3] "rastergrob" "grob" "gDesc"
R
# DAPI背景的细胞分群图
options(repr.plot.height=7, repr.plot.width=15)
ImageSpacePlot(obj=obj, group_by = "Sub_CellType",type="DAPI",size=0.7)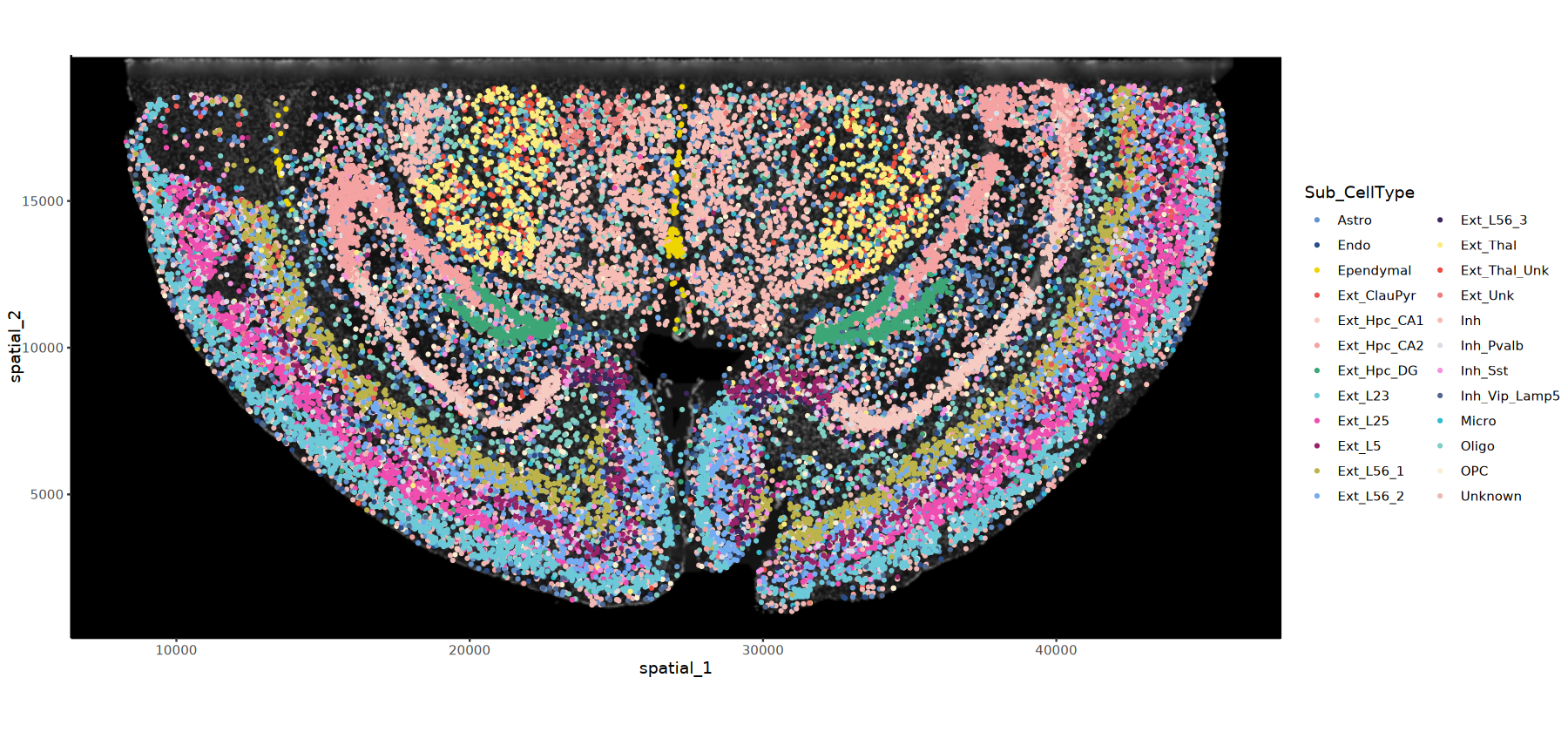
R
# HE背景的细胞分群图
options(repr.plot.height=7.5, repr.plot.width=13)
ImageSpacePlot(obj=obj, group_by = "Sub_CellType",type="HE",size = 0.5)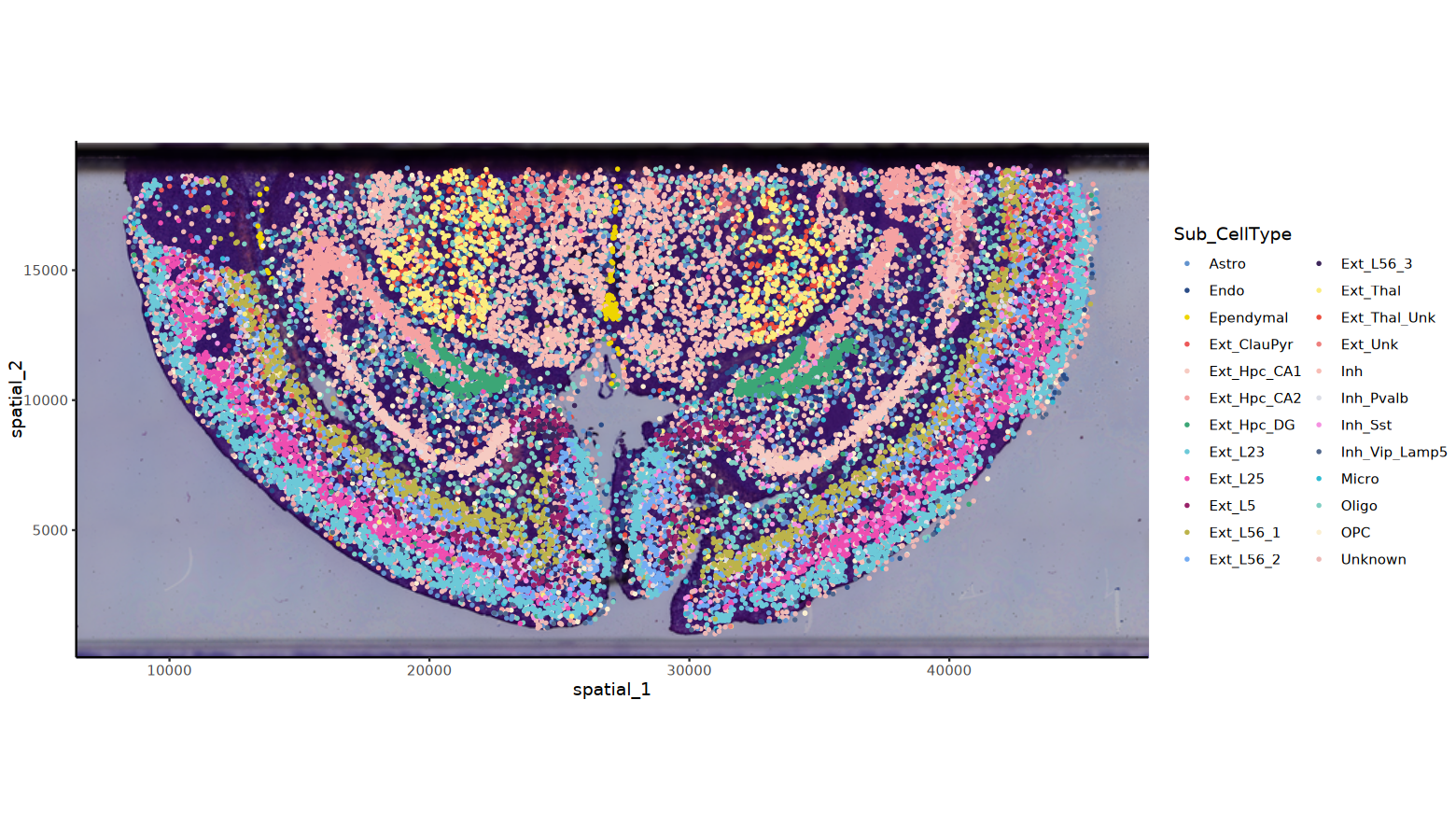
R
# DAPI背景的基因表达图
options(repr.plot.height=7, repr.plot.width=15)
FeatureSpacePlot(obj=obj, feature="Hpca",type="DAPI")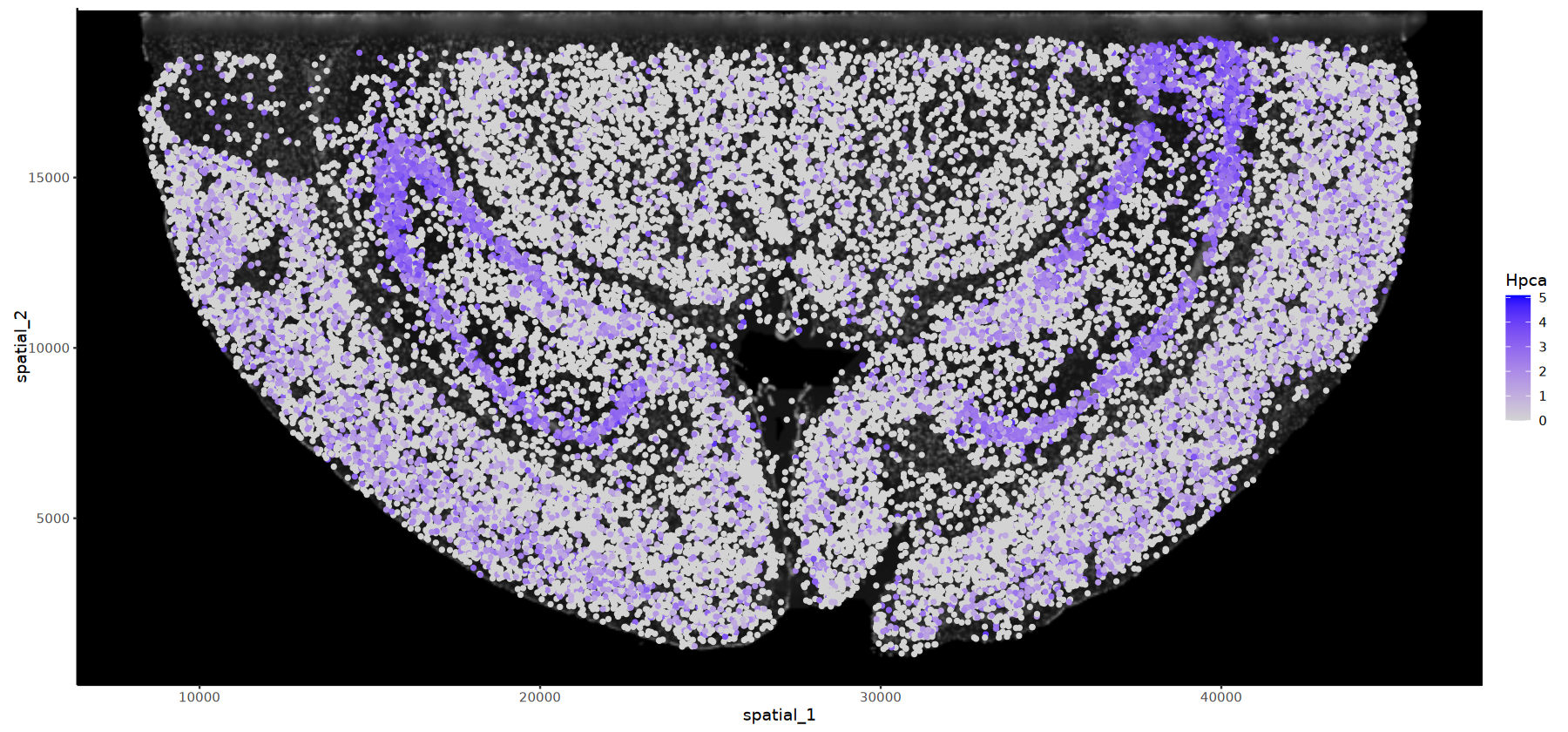
R
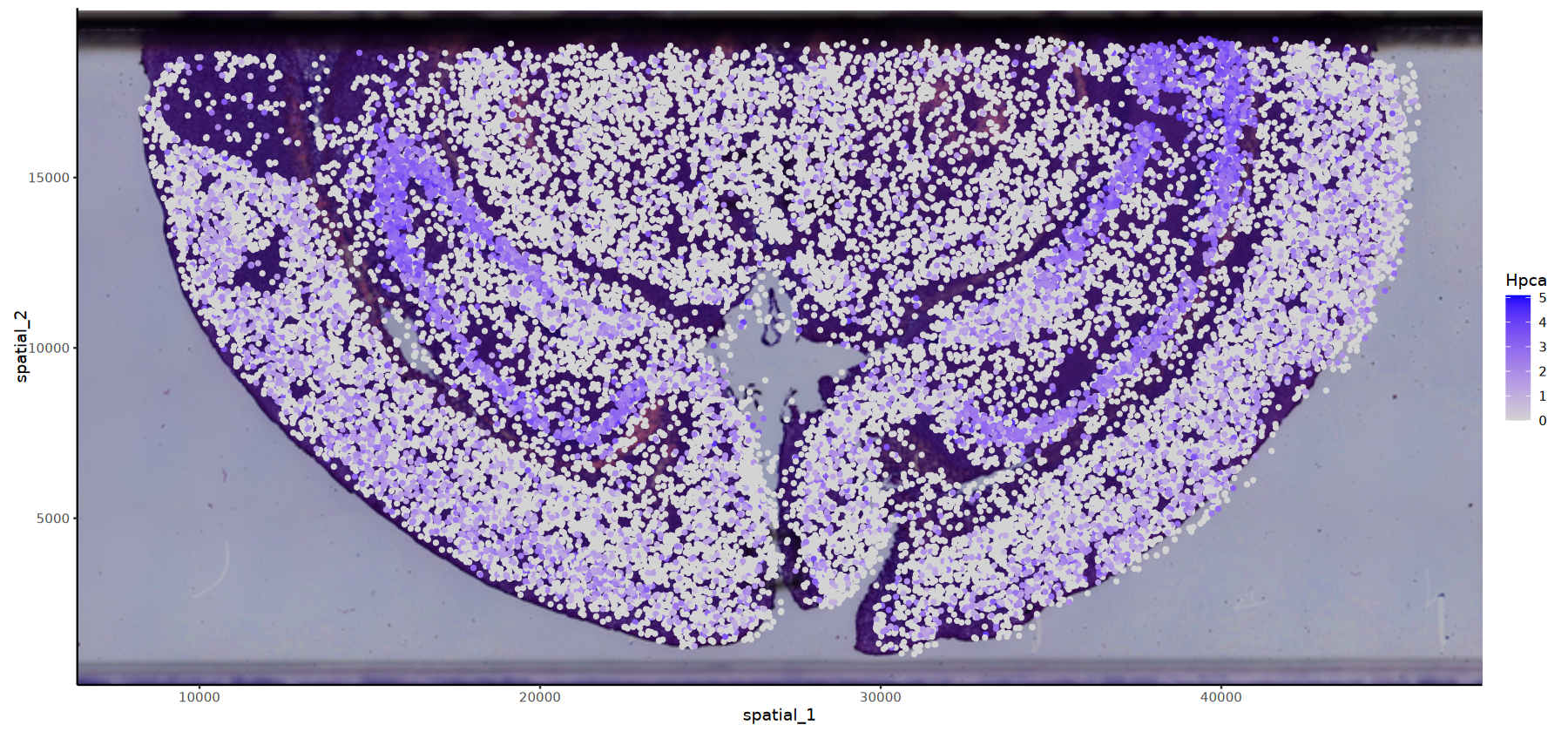
# HE背景的基因表达图
options(repr.plot.height=7, repr.plot.width=15)
FeatureSpacePlot(obj=obj, feature="Hpca",type="HE")