Single-cell Spatial Transcriptomics: Spatial Clustering Analysis (Based on Banksy)
Module Introduction
This module is based on the Banksy algorithm and is designed for spatial clustering and domain segmentation of spatial transcriptomics data.
Banksy is an algorithm that enhances clustering performance by leveraging spatial neighborhood information. It combines gene expression features with spatial microenvironment features to achieve the following core analysis objectives:
- Spatial Domain Identification: Partitions the tissue into biologically meaningful spatial domains (e.g., tumor, stroma, immune infiltration regions) based on gene expression patterns and spatial neighborhood features.
- Denoising and Enhancement: Utilizes neighborhood information to smooth gene expression and reduce technical noise.
- Visualization: Intuitively displays the distribution of different spatial domains.
Comprehensive Analysis: From "Cell Type" to "Niche Function"
Banksy performs spatial clustering by constructing a gene-cell expression matrix and a neighborhood cell-expression matrix using the gene expression matrix and spatial coordinate information. The spatial clustering results (Clusters) represent cells residing in similar microenvironments.
Through Banksy spatial clustering, we can explore the following biological questions:
Who is where? (Spatial Domain + Cell Type) Which cell types are enriched in which specific spatial domains? For instance, do "actively proliferating tumor cells" specifically aggregate in the "hypoxic core domain", while "exhausted T cells" are restricted to the "tumor invasive margin domain"? Such spatial distribution features help reveal the heterogeneity of the cellular microenvironment.
What are they doing there? (Spatially Specific Expression + Cell State) How do the gene expression patterns of cells change within specific spatial domains? Which key functional pathways are enriched in that region? For example, a tumor cell subpopulation located in the "hypoxic core domain" might specifically upregulate genes related to the glycolysis pathway, whereas tumor cells far from this region do not.
How do they interact? (Cell-Cell Communication + Spatial Proximity) Is there active signal crosstalk between spatially adjacent cell populations? Does a ligand expressed in one spatial domain (e.g., fibroblast-enriched region) match a receptor expressed in a directly adjacent spatial domain (e.g., immune cell infiltration region)? By integrating spatial location information, a more biologically meaningful cell-cell communication network map can be constructed.
Disease Progression and Heterogeneity? (Spatial Evolution + Clinical Pathology) Do the spatial clustering results align with the histological structures annotated by pathologists (e.g., normal tissue, precancerous lesions, invasive carcinoma)? Are there significant differences in the composition and distribution of spatial niches between different samples (e.g., pre-treatment vs. post-treatment, or different patients)? Such differences may provide vital clues for the mechanisms of disease progression or treatment response.
💡 Note
This workflow requires a Seurat object (.rds file) as input. The analysis results will be saved as a SpatialExperiment object along with related plots/tables.
Input File Preparation
File Structure Example:
./
├── input.rds # Required: Input Seurat object (must contain RNA assay and spatial coordinates)
└── meta.txt # Optional: External Metadata text fileInput File Details:
Seurat Object (
input.rds):- Expression Matrix: Must contain gene expression data for cells or spatial spots that have undergone preliminary quality control (QC).
- Spatial Coordinates: Must contain physical spatial coordinates (usually stored in
reductions$spatial). The Banksy algorithm relies on these coordinates to compute spatial neighborhood features. - Sample Identifier: The internal metadata of the object should contain a column (e.g.,
Sample) that clearly distinguishes different slices or samples. This is particularly critical in multi-sample integration mode.
External Metadata File (
meta.txt):- If the Seurat object lacks important metadata, or if you need to update sample grouping, clinical information, etc., you can import an external tab-separated text file (e.g.,
.txtor.tsv). - The file should contain a
barcodes(orbarcode) column as row names to accurately match the cells/spatial spots in the Seurat object.
- If the Seurat object lacks important metadata, or if you need to update sample grouping, clinical information, etc., you can import an external tab-separated text file (e.g.,
Data Loading and Preprocessing
This step converts the Seurat object into a SpatialExperiment object required by Banksy and completes basic data processing, including the following stages:
- Load Seurat Object: Read the Seurat object containing spatial coordinates from the
.rdsfile. - Extract Coordinates and Convert Format: Extract spatial coordinates and convert the data into the SpatialExperiment format.
- Data Normalization: Choose between full-gene normalization or highly variable gene extraction based on the dataset size.
# --- Environment Setup ---
# Load core R packages related to spatial transcriptomics analysis and Banksy.
library(Seurat)
library(Banksy)
library(SummarizedExperiment)
library(SpatialExperiment)
library(scuttle)
library(scater)
library(cowplot)
library(ggplot2)
library(tidyverse)
library(futile.logger)
library(future)
library(parallel)
library(doParallel)
library(dplyr)
library(tibble)
library(data.table)
library(harmony)
library(base64enc, quietly = TRUE)
library(reshape2)
library(scales)
library(patchwork)Warning message:
“package ‘SummarizedExperiment’ was built under R version 4.3.2”
Loading required package: MatrixGenerics
Warning message:
“package ‘MatrixGenerics’ was built under R version 4.3.3”
Loading required package: matrixStats
Warning message:
“package ‘matrixStats’ was built under R version 4.3.3”
Attaching package: ‘MatrixGenerics’
The following objects are masked from ‘package:matrixStats’:
colAlls, colAnyNAs, colAnys, colAvgsPerRowSet, colCollapse,
colCounts, colCummaxs, colCummins, colCumprods, colCumsums,
colDiffs, colIQRDiffs, colIQRs, colLogSumExps, colMadDiffs,
colMads, colMaxs, colMeans2, colMedians, colMins, colOrderStats,
colProds, colQuantiles, colRanges, colRanks, colSdDiffs, colSds,
colSums2, colTabulates, colVarDiffs, colVars, colWeightedMads,
colWeightedMeans, colWeightedMedians, colWeightedSds,
colWeightedVars, rowAlls, rowAnyNAs, rowAnys, rowAvgsPerColSet,
rowCollapse, rowCounts, rowCummaxs, rowCummins, rowCumprods,
rowCumsums, rowDiffs, rowIQRDiffs, rowIQRs, rowLogSumExps,
rowMadDiffs, rowMads, rowMaxs, rowMeans2, rowMedians, rowMins,
rowOrderStats, rowProds, rowQuantiles, rowRanges, rowRanks,
rowSdDiffs, rowSds, rowSums2, rowTabulates, rowVarDiffs, rowVars,
rowWeightedMads, rowWeightedMeans, rowWeightedMedians,
rowWeightedSds, rowWeightedVars
Loading required package: GenomicRanges
Warning message:
“package ‘GenomicRanges’ was built under R version 4.3.3”
Loading required package: stats4
Loading required package: BiocGenerics
Warning message:
“package ‘BiocGenerics’ was built under R version 4.3.2”
Attaching package: ‘BiocGenerics’
The following objects are masked from ‘package:stats’:
IQR, mad, sd, var, xtabs
The following objects are masked from ‘package:base’:
anyDuplicated, aperm, append, as.data.frame, basename, cbind,
colnames, dirname, do.call, duplicated, eval, evalq, Filter, Find,
get, grep, grepl, intersect, is.unsorted, lapply, Map, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int,
Position, rank, rbind, Reduce, rownames, sapply, setdiff, sort,
table, tapply, union, unique, unsplit, which.max, which.min
Loading required package: S4Vectors
Warning message:
“package ‘S4Vectors’ was built under R version 4.3.3”
Attaching package: ‘S4Vectors’
The following object is masked from ‘package:utils’:
findMatches
The following objects are masked from ‘package:base’:
expand.grid, I, unname
Loading required package: IRanges
Warning message:
“package ‘IRanges’ was built under R version 4.3.3”
Loading required package: GenomeInfoDb
Warning message:
“package ‘GenomeInfoDb’ was built under R version 4.3.2”
Loading required package: Biobase
Warning message:
“package ‘Biobase’ was built under R version 4.3.3”
Welcome to Bioconductor
Vignettes contain introductory material; view with
'browseVignettes()'. To cite Bioconductor, see
'citation("Biobase")', and for packages 'citation("pkgname")'.
Attaching package: ‘Biobase’
The following object is masked from ‘package:MatrixGenerics’:
rowMedians
The following objects are masked from ‘package:matrixStats’:
anyMissing, rowMedians
Attaching package: ‘SummarizedExperiment’
The following object is masked from ‘package:SeuratObject’:
Assays
The following object is masked from ‘package:Seurat’:
Assays
Loading required package: SingleCellExperiment
Loading required package: ggplot2
Warning message:
“package ‘cowplot’ was built under R version 4.3.3”
Warning message:
“package ‘tibble’ was built under R version 4.3.3”
Warning message:
“package ‘tidyr’ was built under R version 4.3.3”
Warning message:
“package ‘dplyr’ was built under R version 4.3.3”
Warning message:
“package ‘stringr’ was built under R version 4.3.3”
Warning message:
“package ‘lubridate’ was built under R version 4.3.3”
── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✗ lubridate::%within%() masks IRanges::%within%()
✗ dplyr::collapse() masks IRanges::collapse()
✗ dplyr::combine() masks Biobase::combine(), BiocGenerics::combine()
✗ dplyr::count() masks matrixStats::count()
✗ dplyr::desc() masks IRanges::desc()
✗ tidyr::expand() masks S4Vectors::expand()
✗ dplyr::filter() masks stats::filter()
✗ dplyr::first() masks S4Vectors::first()
✗ dplyr::lag() masks stats::lag()
✗ ggplot2::Position() masks BiocGenerics::Position(), base::Position()
✗ purrr::reduce() masks GenomicRanges::reduce(), IRanges::reduce()
✗ dplyr::rename() masks S4Vectors::rename()
✗ lubridate::second() masks S4Vectors::second()
✗ lubridate::second<-() masks S4Vectors::second<-()
✗ dplyr::slice() masks IRanges::slice()
✗ lubridate::stamp() masks cowplot::stamp()
ℹ Use the conflicted package (
Warning message:
“package ‘futile.logger’ was built under R version 4.3.3”
Warning message:
“package ‘future’ was built under R version 4.3.3”
Warning message:
“package ‘doParallel’ was built under R version 4.3.3”
Loading required package: foreach
Warning message:
“package ‘foreach’ was built under R version 4.3.3”
Attaching package: ‘foreach’
The following objects are masked from ‘package:purrr’:
accumulate, when
Loading required package: iterators
Warning message:
“package ‘iterators’ was built under R version 4.3.3”
Warning message:
“package ‘data.table’ was built under R version 4.3.3”
Attaching package: ‘data.table’
The following objects are masked from ‘package:lubridate’:
hour, isoweek, mday, minute, month, quarter, second, wday, week,
yday, year
The following objects are masked from ‘package:dplyr’:
between, first, last
The following object is masked from ‘package:purrr’:
transpose
The following object is masked from ‘package:SummarizedExperiment’:
shift
The following object is masked from ‘package:GenomicRanges’:
shift
The following object is masked from ‘package:IRanges’:
shift
The following objects are masked from ‘package:S4Vectors’:
first, second
Loading required package: Rcpp
Warning message:
“package ‘Rcpp’ was built under R version 4.3.3”
Attaching package: ‘DT’
The following object is masked from ‘package:SeuratObject’:
JS
The following object is masked from ‘package:Seurat’:
JS
Warning message:
“package ‘reshape2’ was built under R version 4.3.3”
Attaching package: ‘reshape2’
The following objects are masked from ‘package:data.table’:
dcast, melt
The following object is masked from ‘package:tidyr’:
smiths
Attaching package: ‘scales’
The following object is masked from ‘package:purrr’:
discard
The following object is masked from ‘package:readr’:
col_factor
# --- Input Parameters Configuration ---
## rds_path: Path to the input Seurat object
rds_path = "./25020230_nao_CS.rds"
## colName: Sample column name in the Seurat object's metadata
colName = "Sample"
## sample_name: Names of the samples to be analyzed.
## - Single-sample mode: Enter a single sample name (e.g., "sample1").
## - Multi-sample integration mode: Enter multiple sample names separated by commas (e.g., "sample1,sample2,sample3"). The system will automatically trigger multi-sample integration and batch correction.
sample_name = "25020230_nao_CS_expression"
## meta_path: Path to external metadata file
meta_path = "./meta.tsv"
## npcs: Number of principal components used during PCA dimensionality reduction
npcs = "30"
## lambda: Spatial weighting parameter, controlling the relative contribution of spatial and expression features. Multiple values are separated by commas.
lambda = "0.6,0.8"
## algo: Clustering algorithm, options include "leiden", "louvain", "kmeans", "mclust"
algo = "leiden"
## resolution: Clustering resolution. Multiple values are separated by commas (applicable for leiden/louvain).
resolution = "0.4,0.8"
## kmeans.centers: Number of kmeans clusters, multiple values separated by commas (applicable for kmeans)
kmeans.centers = "5,10,15,20"
## mclust.G: Number of clusters for mclust. Multiple values are separated by commas. (applicable for mclust)
mclust.G = "5,10,15,20"# Parameter parsing
# Parse and process various input analysis parameters, such as spatial weight (`lambda`), clustering resolution (`resolution`), etc.
if(class(lambda) == 'character'){
lambda = as.numeric(strsplit(as.character(lambda),",")[[1]])
}
if(class(resolution) == 'character'){
resolution = as.numeric(strsplit(as.character(resolution),",")[[1]])
}
if(class(kmeans.centers) == 'character'){
kmeans.centers = as.numeric(strsplit(as.character(kmeans.centers),",")[[1]])
}
if(class(mclust.G) == 'character'){
mclust.G = as.numeric(strsplit(as.character(mclust.G),",")[[1]])
}
npcs = as.numeric(npcs)
if (any(resolution == "")) {
resolution = NULL
}else if (any(kmeans.centers == "")) {
kmeans.centers = NULL
}else if (any(mclust.G == "")) {
mclust.G = NULL
}
if(is.null(resolution)){
rm(resolution)
}else if(is.null(kmeans.centers)){
rm(kmeans.centers)
}else if(is.null(mclust.G)){
rm(mclust.G)
}
sample_name = strsplit(sample_name,",")[[1]]💡 Note
Note on Multi-sample Coordinate Processing: During multi-sample integration, the original physical coordinates of different slices often overlap. Therefore, when extracting coordinate information, this code automatically translates (adds a specific offset to) the coordinate points of different slices based on the sample identifier. This spreads them out flat on the same virtual canvas to prevent erroneous association of cells from different slices when computing the spatial neighborhood network.
# --- Data Loading and Coordinate Processing ---
# Read Seurat object and metadata, extract cell spatial coordinates, and perform coordinate standardization and translation processing based on samples.
obj<-readRDS(rds_path)
meta = read.delim(meta_path)
obj@meta.data = meta
if(colnames(meta)[1]=="barcodes"){
obj@meta.data = column_to_rownames(obj@meta.data,"barcodes")
} else {
obj@meta.data = column_to_rownames(obj@meta.data,"barcode")
}
columns_to_keep <- colnames(obj@meta.data)[!grepl("clust_|HarBsy_", colnames(obj@meta.data))]
obj@meta.data = obj@meta.data[,columns_to_keep]
obj_sample = subset(obj,cells = rownames(obj@meta.data[obj@meta.data[[colName]] %in% sample_name,]))
spatialCoords <- obj_sample@reductions$spatial@cell.embeddings
if(length(sample_name)>1){
spatialCoords = as.data.frame(spatialCoords)
spatialCoords$Sample = obj_sample@meta.data[[colName]]
spatialCoords$sample_id = as.numeric(factor(spatialCoords$Sample, levels = unique(spatialCoords$Sample)))
spatialCoords = spatialCoords[,-which(colnames(spatialCoords) %in% "Sample")]
locs_dt <- data.table(spatialCoords)
colnames(locs_dt) <- c("sdimx", "sdimy", "group")
locs_dt[, sdimx := sdimx - min(sdimx), by = group]
locs_dt$group = as.numeric(locs_dt$group)
global_max <- max(locs_dt$sdimx) * 1.5
locs_dt[, sdimx := sdimx + group * global_max]
locs <- as.matrix(locs_dt[, 1:2])
rownames(locs) <- rownames(obj_sample@meta.data)
} else if(length(sample_name)==1){
colnames(spatialCoords) <- c("sdimx", "sdimy")
}# --- Object Conversion ---
# Convert Seurat object to `SpatialExperiment` object to meet the input format requirements of the Banksy algorithm.
sce <- Seurat::as.SingleCellExperiment(obj_sample)
sample_id <- as.character(unique(obj_sample$Sample))
sample <- gsub(sample_id,pattern = "_expression",replacement = "")
# Convert to SpatialExperiment object, which serves as input for banksy
if(length(sample_name)>1){
spe = SpatialExperiment(
assays = assays(sce),
rowData = rowData(sce),
colData = colData(sce),
spatialCoords = locs
)
} else if(length(sample_name)==1){
spe = SpatialExperiment(
assays = assays(sce),
rowData = rowData(sce),
colData = colData(sce),
sample_id = sample_id,
spatialCoords = spatialCoords,
)
}# --- Data Normalization and Highly Variable Gene Selection ---
# Automatically select processing strategy based on dataset size: use all genes for normalization for small datasets, extract highly variable genes (HVGs) for large datasets for subsequent analysis.
if (dim(spe)[2] <= 50000 & dim(spe)[1] <= 40000) {
# Default is to use all genes. If using all genes, cell count must be within 50000. Normalize the data.
spe <- computeLibraryFactors(spe)
aname <- "normcounts"
assay(spe, aname) <- normalizeCounts(spe, log = FALSE)
} else {
# Use highly variable genes
seu <- as.Seurat(spe, data = NULL)
seu <- FindVariableFeatures(seu, nfeatures = 2000)
# Normalization
scale_factor <- median(colSums(assay(spe, "counts")))
seu <- NormalizeData(seu, scale.factor = scale_factor, normalization.method = "RC")
# Add normalization results to SpatialExperiment object
aname <- "normcounts"
assay(spe, aname) <- GetAssayData(seu)
spe <- spe[VariableFeatures(seu),]
}Spatial Feature Computation and Clustering Core
Banksy accomplishes domain identification through Computing Neighborhood Features, Dimensionality Reduction, and Spatial Clustering. This process includes the following key steps:
- Feature Computation: Enhances gene expression features using spatial neighborhood information.
- Dimensionality Reduction: Performs dimensionality reduction in the augmented feature space using PCA and UMAP.
- Batch Effect Correction: If multiple samples are present, utilizes the Harmony algorithm to perform batch effect correction on the spatial features across the multiple samples.
- Clustering: Executes spatial microenvironment clustering based on the dimensionality reduction results.
# --- Banksy Feature Computation and Dimensionality Reduction ---
# Calculate Banksy enhanced feature matrix based on spatial neighborhood, and perform PCA dimensionality reduction to extract main spatial features.
k_geom <- c(15, 30)
spe <- Banksy::computeBanksy(spe, assay_name = aname, compute_agf = TRUE, k_geom = k_geom)
lambda <- lambda
set.seed(1000)
spe <- Banksy::runBanksyPCA(spe, use_agf = TRUE, lambda = lambda, npcs = npcs)“sparse->dense coercion: allocating vector of size 10.1 GiB”
Computing neighbors...n
Spatial mode is kNN_median
Parameters: k_geom=15
Done
Computing neighbors...n
Spatial mode is kNN_median
Parameters: k_geom=30
Done
Computing harmonic m = 0
Using 15 neighbors
Done
Computing harmonic m = 1
Using 30 neighbors
Centering
Done
Warning message in asMethod(object):
“sparse->dense coercion: allocating vector of size 10.1 GiB”
Warning message in asMethod(object):
“sparse->dense coercion: allocating vector of size 10.1 GiB”
Single-sample and Multi-sample Integrated Clustering:
This tutorial supports both single-sample spatial clustering and multi-sample integrated spatial clustering. The system will automatically determine the analysis mode based on the number of samples (sample_name parameter) you input in the subsequent configuration:
- Single-sample mode: Performs spatial feature enhancement and clustering only on a single tissue slice.
- Multi-sample mode: For multiple tissue slices, after spatial feature computation, it automatically introduces the Harmony algorithm for batch effect correction, followed by joint dimensionality reduction and clustering. This ensures that technical batch differences are eliminated while preserving true spatial biological heterogeneity.
# --- Batch Effect Correction (Multi-sample mode only) ---
if(length(sample_name)>1){
for (i in lambda) {
set.seed(1000)
pca_name <- paste0("PCA_M1_lam", i)
print(paste0("computeBanksy Harmony: ", pca_name))
harmony_embedding <- HarmonyMatrix(
data_mat = reducedDim(spe, pca_name),
meta_data = colData(spe),
vars_use = colName,
do_pca = FALSE,
max.iter.harmony = 20,
verbose = FALSE
)
banksy_name <- paste0("HarBsy_lam", i)
reducedDim(spe, banksy_name) <- harmony_embedding
}
}Differential Processing for Dimensionality Reduction and Clustering in Single/Multi-sample:
- Multi-sample integration mode: Performs UMAP dimensionality reduction and clustering based on Harmony-corrected features (
dimred = 'HarBsy_lam...'). - Single-sample mode: Directly uses uncorrected original Banksy spatial features for subsequent dimensionality reduction and clustering.
# --- Dimensionality Reduction and Spatial Clustering ---
if(length(sample_name)>1){
print("computeBanksy UMAP")
#spe <- Banksy::runBanksyUMAP(spe, use_agf = TRUE, lambda = lambda, npcs = npcs)
for (i in lambda) {
banksy_name <- paste0("HarBsy_lam", i)
spe <- Banksy::runBanksyUMAP(spe, dimred = banksy_name)
}
for (i in lambda) {
banksy_name <- paste0("HarBsy_lam", i)
if(algo == "leiden"|algo == "louvain"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = 'leiden', npcs = npcs, resolution = resolution,dimred = banksy_name)
}else if(algo == "kmeans"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "kmeans",npcs = npcs, kmeans.centers = kmeans.centers,dimred = banksy_name)
}else if(algo == "mclust"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "mclust",npcs = npcs, mclust.G = mclust.G,dimred = banksy_name)
}
}
spe <- Banksy::connectClusters(spe)
} else if(length(sample_name)==1){
spe <- Banksy::runBanksyUMAP(spe, use_agf = TRUE, lambda = lambda, npcs = npcs)
if(algo == "leiden"|algo == "louvain"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = 'leiden', npcs = npcs, resolution = resolution)
}else if(algo == "kmeans"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "kmeans",npcs = npcs, kmeans.centers = kmeans.centers)
}else if(algo == "mclust"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "mclust",npcs = npcs, mclust.G = mclust.G)
}
spe <- Banksy::connectClusters(spe)
}clust_M1_lam0.8_k50_res0.8 --> clust_M1_lam0.6_k50_res0.4
clust_M1_lam0.6_k50_res0.8 --> clust_M1_lam0.8_k50_res0.8
colData(spe) <- cbind(colData(spe), spatialCoords(spe))
data = as.data.frame(colData(spe))
color_palette_24 = c("#E41A1C", "#377EB8", "#4DAF4A", "#984EA3", "#FF7F00",
"#FFFF33", "#A65628", "#F781BF", "#999999", "#66C2A5",
"#FC8D62", "#8DA0CB", "#E78AC3", "#A6D854", "#FFD92F",
"#E5C494", "#B3B3B3", "#6A3D9A", "#FF4500", "#2E8B57",
"#9467BD", "#8C564B", "#E377C2", "#7F7F7F", "#BCBD22",
"#17BECF", "#A6761D", "#1B9E77", "#D95F02", "#7570B3",
"#E7298A", "#66A61E", "#E6AB02", "#A6761D", "#666666",
"#A6CEE3", "#1F78B4", "#B2DF8A", "#33A02C", "#FB9A99",
"#E31A1C", "#FDBF6F", "#FF7F00", "#CAB2D6", "#6A3D9A",
"#FFFF99", "#B15928", "#BDC3C7", "#2C3E50", "#E67E22")
cnames <- grep("^clust", colnames(colData(spe)), value = TRUE)Spatial Clustering Visualization
Figure Legend: The figure below shows the spatial clustering map of the tissue slice.
- Axes: Represent the physical spatial coordinates of the tissue slice, mapping the cells' positions within the real tissue.
- Color Encoding: Different colors distinguish different spatial clustering groups (Clusters). Each cluster can be understood as a tissue microenvironment (or niche) with specific molecular features and spatial proximity.
- Title Parameters Explanation:
- lam: Represents the spatial weighting parameter (λ), determining the mixing ratio between the cell's own expression and the surrounding microenvironment features. A larger λ value means the clustering result is more inclined to consider spatial neighborhood information, making physically proximal cells more likely to be clustered together, forming smoother and more continuous spatial domains; a smaller λ value relies more heavily on the cell's own transcriptomic features.
- Clustering Parameter: Depending on the chosen algorithm, the numerical value after
resindicates the clustering resolution for the leiden/louvain algorithm; the numerical value afterkmeans/mclustindicates the preset number of clusters.💡 Analysis Advice & Optimization: The granularity of spatial clustering is jointly controlled by the spatial weight (λ) and the clustering parameter (
res/kmeans/mclust). If the current clustering result is too fragmented, you can manually merge adjacent or similar regions based on anatomical structures in the subsequent cloud platform interactive interface.
options(repr.plot.height=8, repr.plot.width=16)
for(i in 1:length(sample_name)){
a = data[data$Sample %in% sample_name[i],]
for(j in 1:length(cnames)){
# Dynamically calculate number of legend columns
num_clusters <- length(unique(a[[cnames[j]]])) # Get the number of categories for the current cluster variable
if(num_clusters <= 10){
ncol_legend <- 1
} else if(num_clusters <= 20){
ncol_legend <- 2
} else if(num_clusters <= 30){
ncol_legend <- 3
} else {
ncol_legend <- 4 # Use 4 columns when exceeding 30
}
plots = ggplot(a, aes(sdimx, sdimy))+
geom_point(aes_string(color=cnames[j], fill = cnames[j]), size = 1.2)+
coord_equal()+
theme_classic()+
theme(plot.background = element_rect(fill = "black", color = NA),
panel.background = element_rect(fill = "black", color = NA),
legend.background = element_rect(fill = "black", color = NA),
legend.key = element_rect(fill = "black", color = NA),
text = element_text(color = "white", size = 16),
axis.text = element_text(color = "white", size = 14),
axis.title = element_text(color = "white", size = 18),
axis.line = element_line(color = "white"),
axis.ticks = element_line(color = "white"),
legend.title = element_text(color = "white", size = 16),
legend.text = element_text(color = "white", size = 14),
legend.position = "right",
legend.spacing = unit(0.5, "cm"),
legend.key.size = unit(1, "cm")) +
scale_color_manual(values = color_palette_24)+
xlab("spatial_1")+
ylab("spatial_2")+
guides(color = guide_legend(override.aes = list(size = 5),
ncol = ncol_legend)) # Use dynamically calculated number of columns
print(plots)
}
}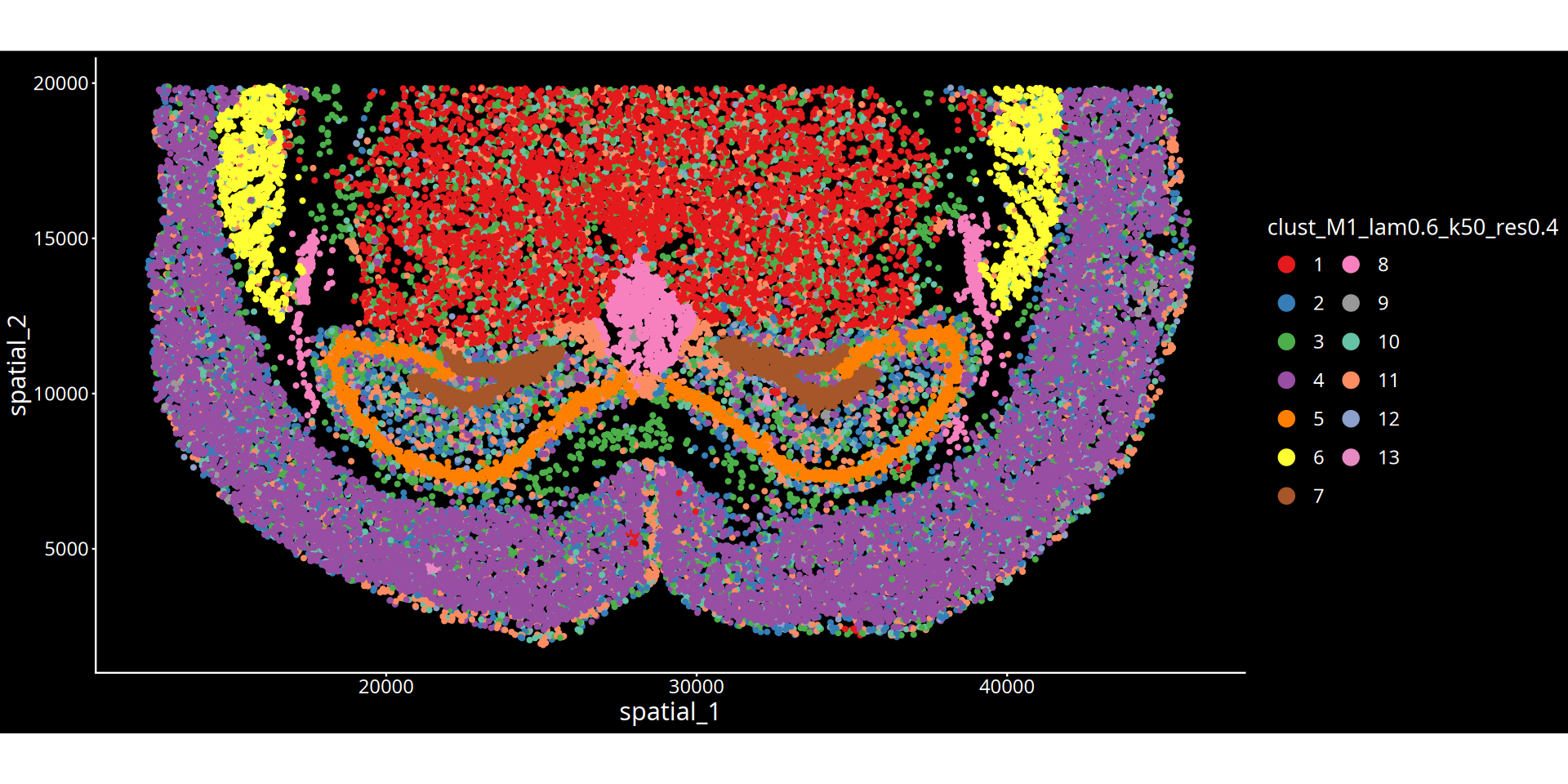
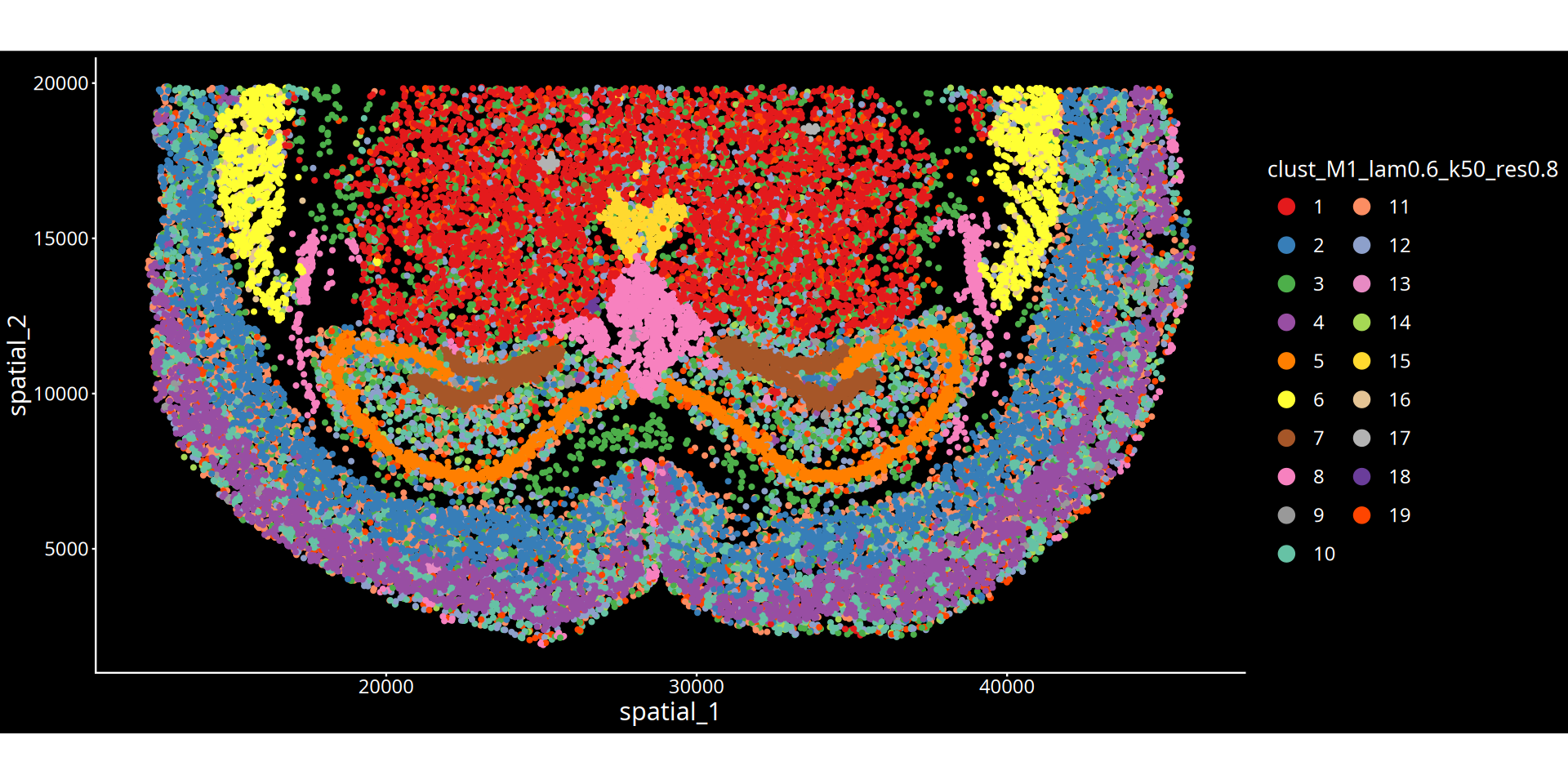
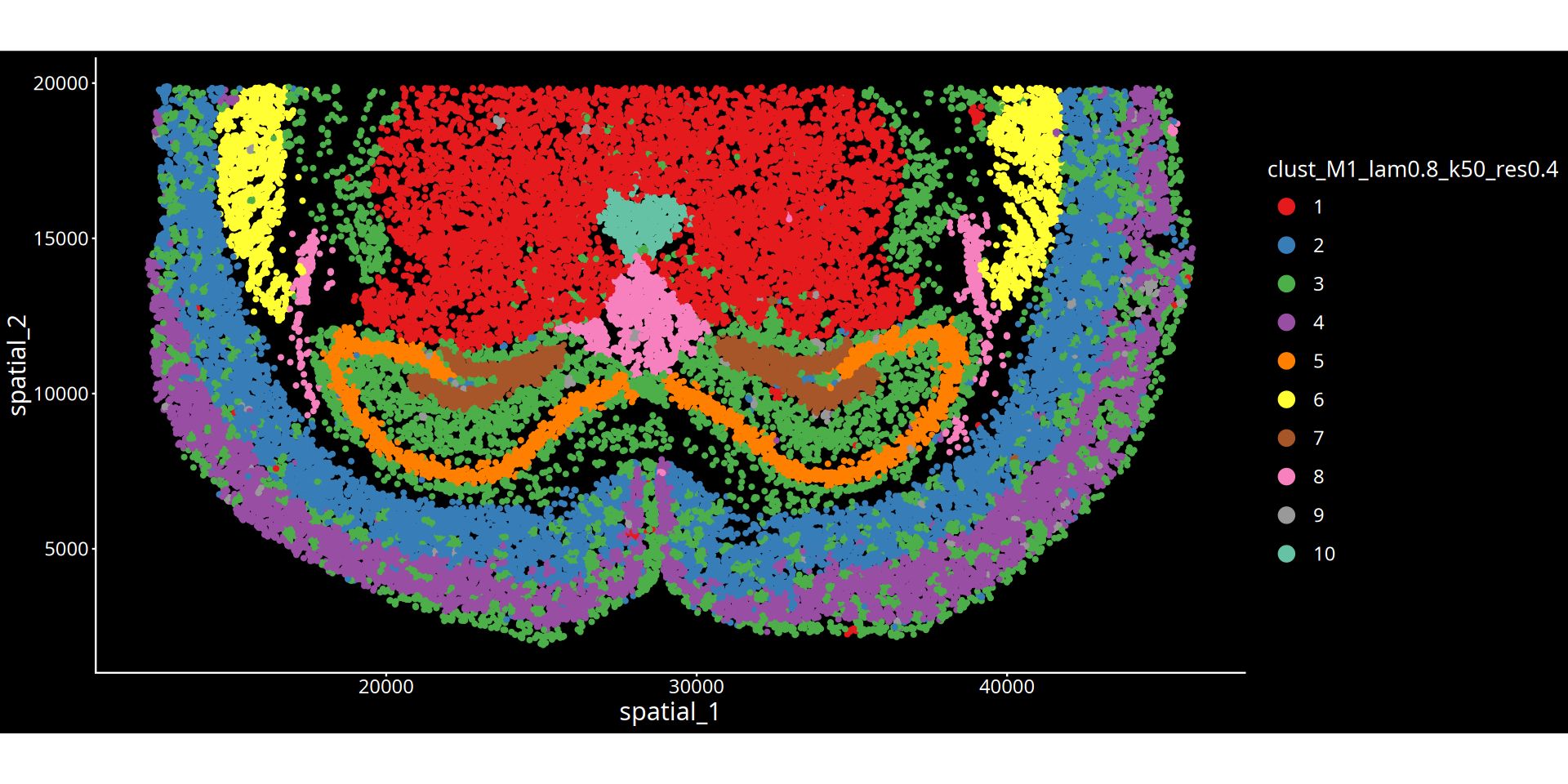
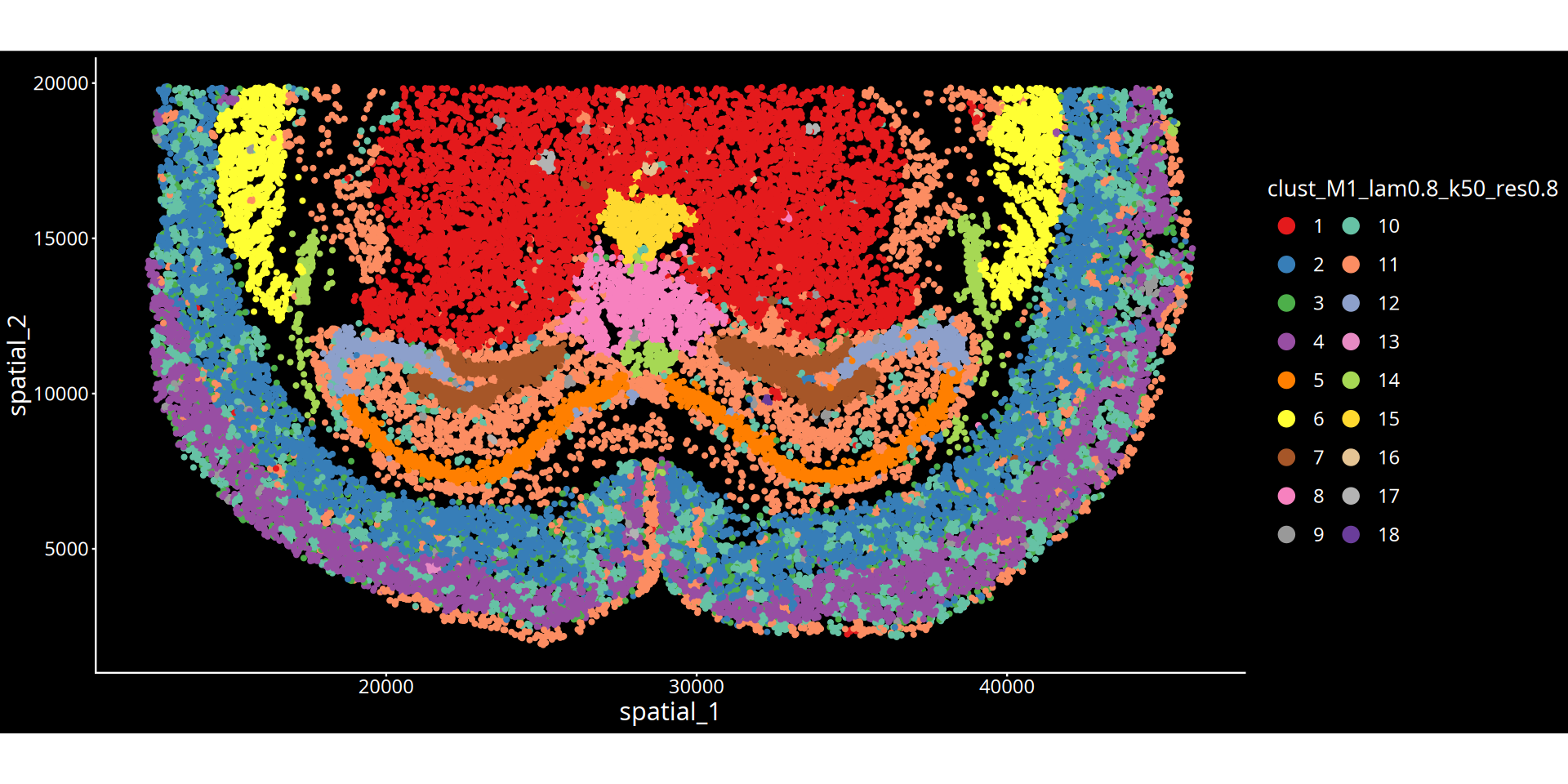
Figure Legend: The figure below displays the UMAP plot of cells in the dimension-reduced space.
- Axes: The two-dimensional coordinates after UMAP dimensionality reduction (UMAP_1 and UMAP_2), preserving the topological structure of the high-dimensional feature space.
- Color Encoding: Spatial clustering result (Cluster).
# Set graphic parameters
options(repr.plot.height=16, repr.plot.width=16)
# Initialize list to store all plots
plot_list <- list()
plot_counter <- 1
# Execute different processing based on sample quantity
if(length(sample_name) > 1){
# Multi-sample scenario
if(algo == "leiden" | algo == "louvain"){
for(i in 1:length(lambda)){
# Get UMAP dimensionality reduction data
b = reducedDim(spe, paste0("UMAP_HarBsy_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
paste0("UMAP_HarBsy_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(resolution)){
# Get clustering column names
cluster_col <- paste0("clust_HarBsy_lam", lambda[i], "_k50_res", resolution[j])
# Calculate number of clusters and set legend columns
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
# Create plot
plots = ggplot(data, aes_string(x = paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
y = paste0("UMAP_HarBsy_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.x = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", res=", resolution[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend),
fill = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
# Store plot
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "kmeans"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_HarBsy_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
paste0("UMAP_HarBsy_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(kmeans.centers)){
cluster_col <- paste0("clust_HarBsy_lam", lambda[i], "_k50_kmeans", kmeans.centers[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
y = paste0("UMAP_HarBsy_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.x = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", kmeans=", kmeans.centers[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend),
fill = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "mclust"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_HarBsy_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
paste0("UMAP_HarBsy_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(mclust.G)){
cluster_col <- paste0("clust_HarBsy_lam", lambda[i], "_k50_mclust", mclust.G[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
y = paste0("UMAP_HarBsy_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.x = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", mclust G=", mclust.G[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend),
fill = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
}
} else if(length(sample_name) == 1){
# Single-sample scenario
if(algo == "leiden" | algo == "louvain"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_M1_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_M1_lam", lambda[i], "_1"),
paste0("UMAP_M1_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(resolution)){
cluster_col <- paste0("clust_M1_lam", lambda[i], "_k50_res", resolution[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_M1_lam", lambda[i], "_1"),
y = paste0("UMAP_M1_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.y = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", res=", resolution[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "kmeans"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_M1_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_M1_lam", lambda[i], "_1"),
paste0("UMAP_M1_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(kmeans.centers)){
cluster_col <- paste0("clust_M1_lam", lambda[i], "_k50_kmeans", kmeans.centers[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_M1_lam", lambda[i], "_1"),
y = paste0("UMAP_M1_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.y = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", kmeans=", kmeans.centers[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "mclust"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_M1_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_M1_lam", lambda[i], "_1"),
paste0("UMAP_M1_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(mclust.G)){
cluster_col <- paste0("clust_M1_lam", lambda[i], "_k50_mclust", mclust.G[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_M1_lam", lambda[i], "_1"),
y = paste0("UMAP_M1_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.y = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", mclust G=", mclust.G[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
}
}
# Combine all plots
# Calculate number of plots displayed per row (can be adjusted as needed)
ncol_layout <- 2 # Display 2 plots per row
nrow_layout <- ceiling(length(plot_list) / ncol_layout)
# Use wrap_plots to combine plots
combined_plot <- wrap_plots(plot_list, ncol = ncol_layout, byrow = TRUE) +
plot_annotation(
title = paste0("Clustering Results - ", toupper(algo)),
theme = theme(plot.title = element_text(size = 16, hjust = 0.5, face = "bold"))
)
# Display combined plots
print(combined_plot)“annotation$theme is not a valid theme.
Please use \`theme()\` to construct themes.”
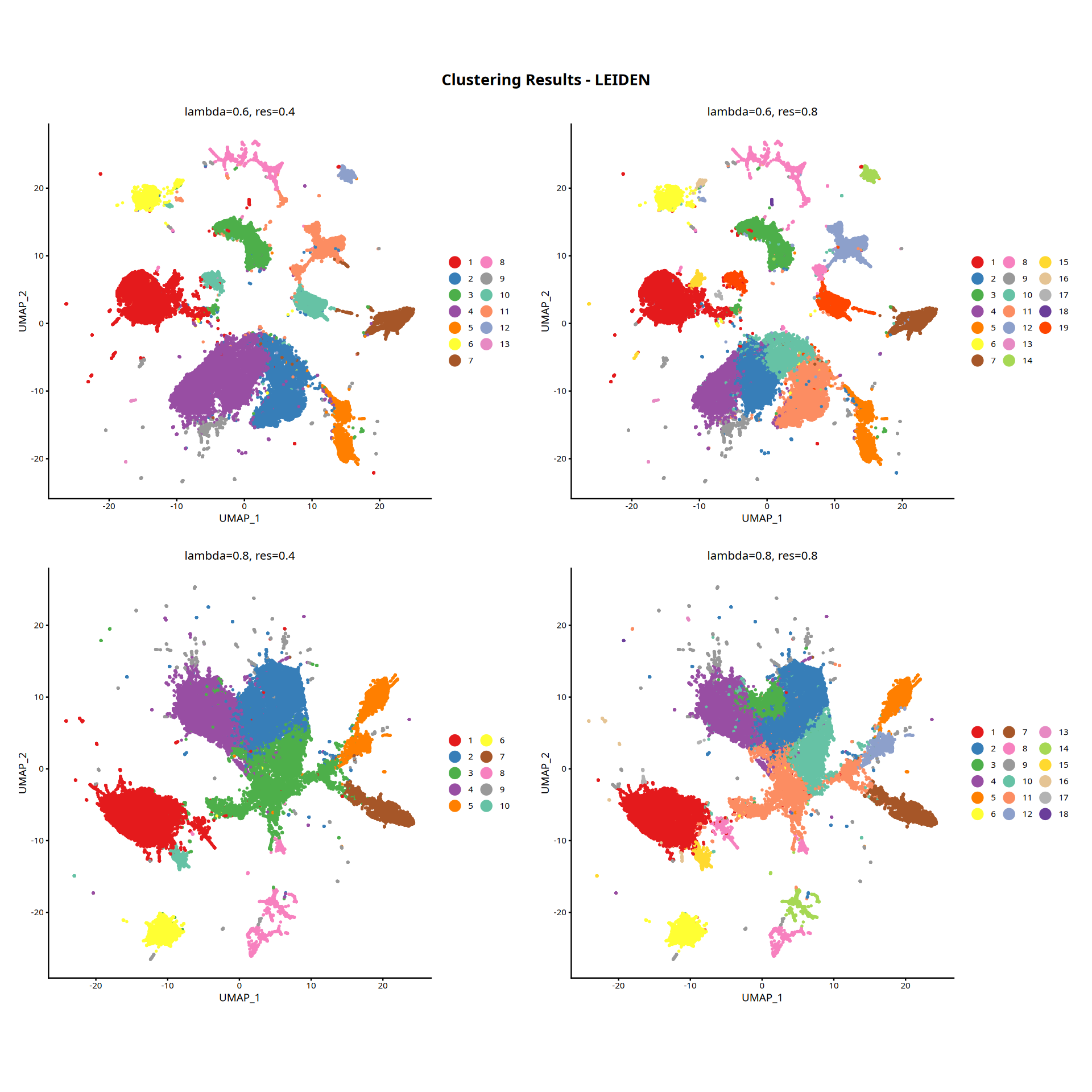
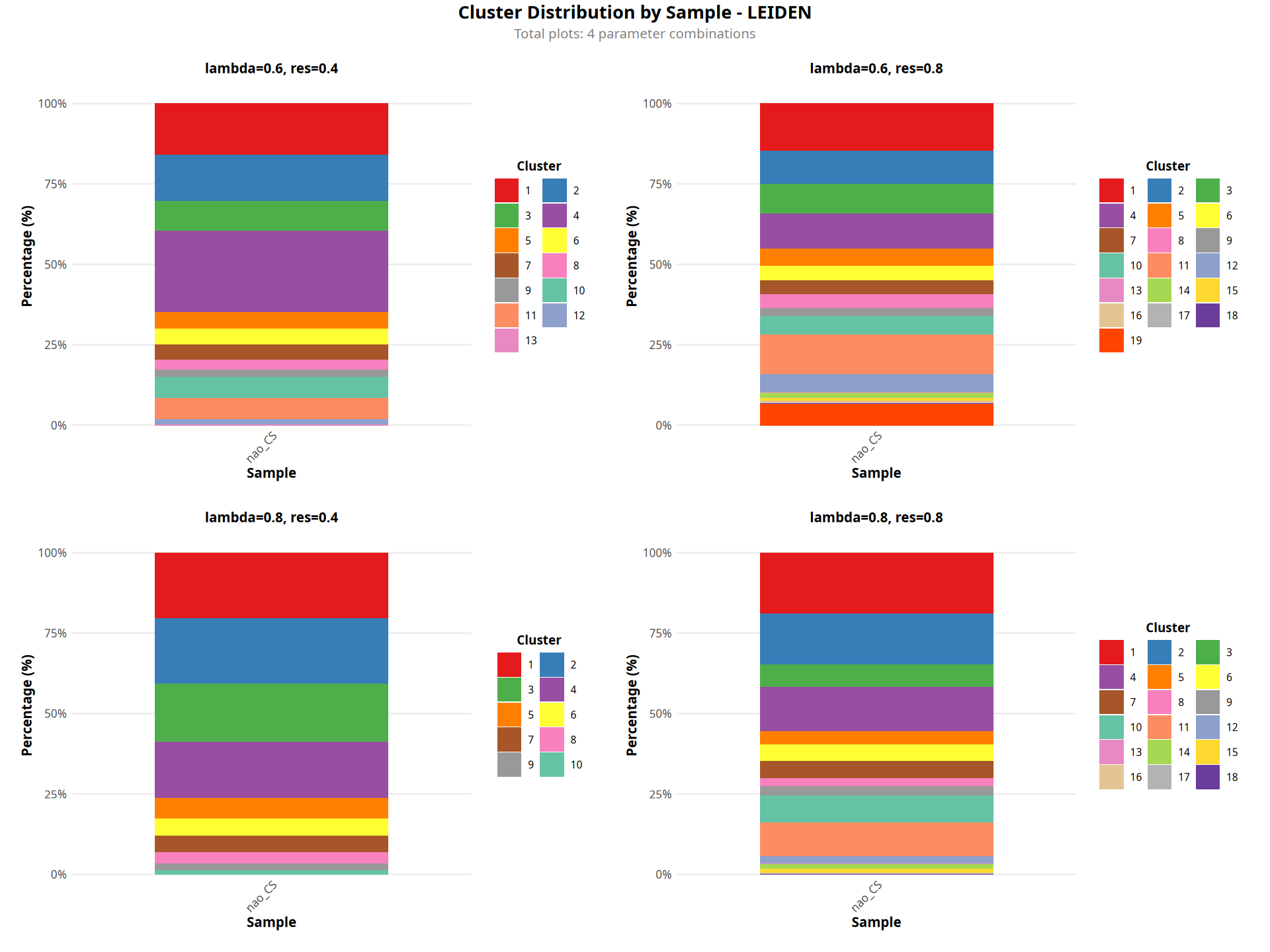
Figure Legend: The figure below shows a stacked bar chart of cluster proportions for each sample.
- X-axis: Sample name.
- Y-axis: Percentage, indicating the proportion of cells for each cluster within the sample.
- Color Encoding: Spatial clustering result (Cluster).
- Interpretation: Used to compare differences in the composition of spatial domains across different samples.
# Load necessary packages
library(ggplot2)
library(reshape2)
library(dplyr)
library(stringr)
library(scales)
library(patchwork)
# Initialize list to store all plots
plot_list <- list()
plot_counter <- 1
# Set graphic parameters
options(repr.plot.height=12, repr.plot.width=16)
for(i in 1:length(cnames)){
# Extract parameters
lam = as.numeric(gsub(".*lam([0-9.]+)_.*", "\\1", cnames[i]))
if(algo == "leiden" | algo == "louvain"){
res = as.numeric(gsub(".*res([0-9.]+).*", "\\1", cnames[i]))
param_label <- paste0("lambda=", lam, ", res=", res)
} else if(algo == "kmeans"){
km = as.numeric(gsub(".*kmeans([0-9]+).*", "\\1", cnames[i]))
param_label <- paste0("lambda=", lam, ", kmeans=", km)
} else if(algo == "mclust"){
g = as.numeric(gsub(".*mclust([0-9]+).*", "\\1", cnames[i]))
param_label <- paste0("lambda=", lam, ", mclust G=", g)
}
# Create basic data from table command
tab <- table(data[[colName]], data[[cnames[i]]])
# Convert to dataframe
df <- as.data.frame.matrix(tab)
df$Sample <- rownames(df)
# Convert to long format
df_long <- melt(df, id.vars = "Sample", variable.name = "Cluster", value.name = "Count")
df_long$Sample = str_replace_all(df_long$Sample, c("^\\d+_" = "", "_expression.*$" = ""))
# Calculate percentage and group by sample
df_long <- df_long %>%
group_by(Sample) %>%
mutate(
Percentage = Count / sum(Count) * 100,
Percentage_label = sprintf("%.1f%%", Percentage)
) %>%
filter(Percentage > 0) %>%
arrange(Sample, desc(Percentage))
# Get number of clusters and automatically set legend columns
n_clusters <- length(unique(df_long$Cluster))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
# Draw plot
p <- ggplot(df_long, aes(x = Sample, y = Percentage, fill = Cluster)) +
geom_bar(stat = "identity", position = "stack", width = 0.7) +
scale_fill_manual(values = color_palette_24[1:n_clusters]) +
labs(
title = param_label,
x = "Sample",
y = "Percentage (%)"
) +
scale_y_continuous(
labels = percent_format(scale = 1),
expand = expansion(mult = c(0, 0.05))
) +
theme_minimal() +
theme(
# X-axis label
axis.text.x = element_text(angle = 45, hjust = 1, size = 10),
axis.title.x = element_text(size = 12, face = "bold"),
# Y-axis label
axis.text.y = element_text(size = 10),
axis.title.y = element_text(size = 12, face = "bold"),
# Title
plot.title = element_text(size = 12, face = "bold", hjust = 0.5, margin = margin(b = 10)),
# Legend settings
legend.position = "right",
legend.title = element_text(size = 11, face = "bold"),
legend.text = element_text(size = 9),
legend.key.size = unit(0.8, "cm"),
legend.spacing.y = unit(0.2, "cm"),
legend.margin = margin(t = 10, r = 10, b = 10, l = 10),
# Grid lines
panel.grid.major.x = element_blank(),
panel.grid.minor = element_blank(),
# Background and margins
panel.background = element_rect(fill = "white", color = NA),
plot.background = element_rect(fill = "white", color = NA),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm")
) +
guides(
fill = guide_legend(
ncol = ncol_legend,
byrow = TRUE,
title = "Cluster",
title.position = "top",
title.hjust = 0.5,
label.position = "right",
label.hjust = 0
)
)
# Store plot
plot_list[[plot_counter]] <- p
plot_counter <- plot_counter + 1
}
# Combine all plots
# Calculate layout (display 2-3 plots per row)
ncol_layout <- ifelse(length(plot_list) >= 6, 3, 2)
nrow_layout <- ceiling(length(plot_list) / ncol_layout)
# Use wrap_plots to combine plots
combined_plot <- wrap_plots(plot_list, ncol = ncol_layout, byrow = TRUE) +
plot_annotation(
title = paste0("Cluster Distribution by Sample - ", toupper(algo)),
subtitle = paste0("Total plots: ", length(plot_list), " parameter combinations"),
theme = theme(
plot.title = element_text(size = 16, hjust = 0.5, face = "bold"),
plot.subtitle = element_text(size = 12, hjust = 0.5, color = "gray50")
)
)
# Display combined plots
print(combined_plot)“annotation$theme is not a valid theme.
Please use \`theme()\` to construct themes.”