单细胞空间转录组: 空间聚类分析(基于 Banksy)
模块简介
本模块基于 Banksy 算法,用于对空间转录组数据进行空间聚类和区域分割 (Domain Segmentation)。
Banksy 是一种利用空间邻域信息增强聚类效果的算法,它结合了基因表达特征和空间微环境特征,实现以下核心分析目标:
- 空间区域识别:根据基因表达模式和空间邻域特征,将组织划分为具有生物学意义的空间区域(如肿瘤区、基质区、免疫浸润区)。
- 去噪与增强:利用邻域信息平滑基因表达,减少技术噪声。
- 可视化:直观展示不同空间区域的分布。
综合分析:从“细胞类型”到“生态位功能”
Banksy 通过基因表达矩阵和空间位置坐标信息,构建基因-细胞表达矩阵与邻域细胞-表达矩阵进行空间聚类,空间聚类结果(Cluster)代表着细胞处在相似的微环境。
通过 Banksy 空间聚类,我们可以探讨以下生物学问题:
谁在哪里?(空间域 + 细胞类型) 哪些细胞类型富集在哪个特定的空间域中?比如,“增殖活跃的肿瘤细胞”是否特异性地聚集在“缺氧核心域”,而“耗竭状态的 T 细胞”是否被限制在“肿瘤浸润边缘域”?这种空间分布特征有助于揭示细胞微环境的异质性。
它们在那儿做什么?(空间特异性表达 + 细胞状态) 在特定空间域中,细胞的基因表达模式发生了怎样的改变?该区域富集了哪些关键的功能通路?例如,位于“缺氧核心域”的肿瘤细胞亚群可能特异性地上调了糖酵解途径相关基因,而远离该区域的肿瘤细胞则不然。
它们如何互动?(细胞间通讯 + 空间邻近性) 空间上相邻的细胞群之间是否存在活跃的信号交流?一个空间域(如成纤维细胞富集区)中表达的配体,是否与紧邻的另一个空间域(如免疫细胞浸润区)中表达的受体相匹配?通过结合空间位置信息,可以构建出更具生物学意义的细胞间通讯网络图。
疾病演进与异质性?(空间演化 + 临床病理) 空间聚类结果是否与病理学家标注的组织学结构(如正常组织、癌前病变、浸润癌)相吻合?不同样本(如治疗前 vs 治疗后,或不同患者)之间空间生态位的组成和分布是否存在显著差异?这种差异可能为疾病的进展机制或治疗响应提供重要线索。
💡 Note
本流程输入为 Seurat 对象(.rds 文件),分析结果将保存为 SpatialExperiment 对象及相关图片/表格。
输入文件准备
文件结构示例:
./
├── input.rds # 必需:输入的 Seurat 对象(需包含 RNA assay 和 spatial 空间坐标信息)
└── meta.txt # 可选:外部的 Metadata 文本文件输入文件详细说明:
Seurat 对象 (
input.rds):- 表达矩阵:需包含经过初步质控的细胞/空间点基因表达数据。
- 空间坐标信息:必须包含空间物理坐标(通常存储在
reductions$spatial),Banksy 算法需要依赖此坐标来计算空间邻域特征。 - 样本标识:对象内部的 metadata 中应包含明确区分不同切片或样本的列(如
Sample),这在多样本整合模式下尤为关键。
外部 Metadata 文件 (
meta.txt):- 若 Seurat 对象中缺失部分重要的元数据,或需要更新样本分组、临床信息等,可通过外部制表符分隔的文本文件(如
.txt或.tsv)导入。 - 文件应包含
barcodes(或barcode)列作为行名,以便与 Seurat 对象中的细胞/空间点准确匹配。
- 若 Seurat 对象中缺失部分重要的元数据,或需要更新样本分组、临床信息等,可通过外部制表符分隔的文本文件(如
数据加载与预处理
本步骤将 Seurat 对象转换为 Banksy 所需的 SpatialExperiment 对象并完成基础数据处理,包含以下环节:
- 加载 Seurat 对象:从
.rds读入带有空间坐标信息的 Seurat 对象。 - 提取坐标并转换格式:提取空间坐标,转换为 SpatialExperiment 格式。
- 数据归一化:根据数据集大小选择全基因归一化或高变基因提取。
# --- 环境准备 ---
# 加载空间组学分析与 Banksy 相关的核心 R 包。
library(Seurat)
library(Banksy)
library(SummarizedExperiment)
library(SpatialExperiment)
library(scuttle)
library(scater)
library(cowplot)
library(ggplot2)
library(tidyverse)
library(futile.logger)
library(future)
library(parallel)
library(doParallel)
library(dplyr)
library(tibble)
library(data.table)
library(harmony)
library(base64enc, quietly = TRUE)
library(reshape2)
library(scales)
library(patchwork)Warning message:
“package ‘SummarizedExperiment’ was built under R version 4.3.2”
Loading required package: MatrixGenerics
Warning message:
“package ‘MatrixGenerics’ was built under R version 4.3.3”
Loading required package: matrixStats
Warning message:
“package ‘matrixStats’ was built under R version 4.3.3”
Attaching package: ‘MatrixGenerics’
The following objects are masked from ‘package:matrixStats’:
colAlls, colAnyNAs, colAnys, colAvgsPerRowSet, colCollapse,
colCounts, colCummaxs, colCummins, colCumprods, colCumsums,
colDiffs, colIQRDiffs, colIQRs, colLogSumExps, colMadDiffs,
colMads, colMaxs, colMeans2, colMedians, colMins, colOrderStats,
colProds, colQuantiles, colRanges, colRanks, colSdDiffs, colSds,
colSums2, colTabulates, colVarDiffs, colVars, colWeightedMads,
colWeightedMeans, colWeightedMedians, colWeightedSds,
colWeightedVars, rowAlls, rowAnyNAs, rowAnys, rowAvgsPerColSet,
rowCollapse, rowCounts, rowCummaxs, rowCummins, rowCumprods,
rowCumsums, rowDiffs, rowIQRDiffs, rowIQRs, rowLogSumExps,
rowMadDiffs, rowMads, rowMaxs, rowMeans2, rowMedians, rowMins,
rowOrderStats, rowProds, rowQuantiles, rowRanges, rowRanks,
rowSdDiffs, rowSds, rowSums2, rowTabulates, rowVarDiffs, rowVars,
rowWeightedMads, rowWeightedMeans, rowWeightedMedians,
rowWeightedSds, rowWeightedVars
Loading required package: GenomicRanges
Warning message:
“package ‘GenomicRanges’ was built under R version 4.3.3”
Loading required package: stats4
Loading required package: BiocGenerics
Warning message:
“package ‘BiocGenerics’ was built under R version 4.3.2”
Attaching package: ‘BiocGenerics’
The following objects are masked from ‘package:stats’:
IQR, mad, sd, var, xtabs
The following objects are masked from ‘package:base’:
anyDuplicated, aperm, append, as.data.frame, basename, cbind,
colnames, dirname, do.call, duplicated, eval, evalq, Filter, Find,
get, grep, grepl, intersect, is.unsorted, lapply, Map, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int,
Position, rank, rbind, Reduce, rownames, sapply, setdiff, sort,
table, tapply, union, unique, unsplit, which.max, which.min
Loading required package: S4Vectors
Warning message:
“package ‘S4Vectors’ was built under R version 4.3.3”
Attaching package: ‘S4Vectors’
The following object is masked from ‘package:utils’:
findMatches
The following objects are masked from ‘package:base’:
expand.grid, I, unname
Loading required package: IRanges
Warning message:
“package ‘IRanges’ was built under R version 4.3.3”
Loading required package: GenomeInfoDb
Warning message:
“package ‘GenomeInfoDb’ was built under R version 4.3.2”
Loading required package: Biobase
Warning message:
“package ‘Biobase’ was built under R version 4.3.3”
Welcome to Bioconductor
Vignettes contain introductory material; view with
'browseVignettes()'. To cite Bioconductor, see
'citation("Biobase")', and for packages 'citation("pkgname")'.
Attaching package: ‘Biobase’
The following object is masked from ‘package:MatrixGenerics’:
rowMedians
The following objects are masked from ‘package:matrixStats’:
anyMissing, rowMedians
Attaching package: ‘SummarizedExperiment’
The following object is masked from ‘package:SeuratObject’:
Assays
The following object is masked from ‘package:Seurat’:
Assays
Loading required package: SingleCellExperiment
Loading required package: ggplot2
Warning message:
“package ‘cowplot’ was built under R version 4.3.3”
Warning message:
“package ‘tibble’ was built under R version 4.3.3”
Warning message:
“package ‘tidyr’ was built under R version 4.3.3”
Warning message:
“package ‘dplyr’ was built under R version 4.3.3”
Warning message:
“package ‘stringr’ was built under R version 4.3.3”
Warning message:
“package ‘lubridate’ was built under R version 4.3.3”
── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✗ lubridate::%within%() masks IRanges::%within%()
✗ dplyr::collapse() masks IRanges::collapse()
✗ dplyr::combine() masks Biobase::combine(), BiocGenerics::combine()
✗ dplyr::count() masks matrixStats::count()
✗ dplyr::desc() masks IRanges::desc()
✗ tidyr::expand() masks S4Vectors::expand()
✗ dplyr::filter() masks stats::filter()
✗ dplyr::first() masks S4Vectors::first()
✗ dplyr::lag() masks stats::lag()
✗ ggplot2::Position() masks BiocGenerics::Position(), base::Position()
✗ purrr::reduce() masks GenomicRanges::reduce(), IRanges::reduce()
✗ dplyr::rename() masks S4Vectors::rename()
✗ lubridate::second() masks S4Vectors::second()
✗ lubridate::second<-() masks S4Vectors::second<-()
✗ dplyr::slice() masks IRanges::slice()
✗ lubridate::stamp() masks cowplot::stamp()
ℹ Use the conflicted package (
Warning message:
“package ‘futile.logger’ was built under R version 4.3.3”
Warning message:
“package ‘future’ was built under R version 4.3.3”
Warning message:
“package ‘doParallel’ was built under R version 4.3.3”
Loading required package: foreach
Warning message:
“package ‘foreach’ was built under R version 4.3.3”
Attaching package: ‘foreach’
The following objects are masked from ‘package:purrr’:
accumulate, when
Loading required package: iterators
Warning message:
“package ‘iterators’ was built under R version 4.3.3”
Warning message:
“package ‘data.table’ was built under R version 4.3.3”
Attaching package: ‘data.table’
The following objects are masked from ‘package:lubridate’:
hour, isoweek, mday, minute, month, quarter, second, wday, week,
yday, year
The following objects are masked from ‘package:dplyr’:
between, first, last
The following object is masked from ‘package:purrr’:
transpose
The following object is masked from ‘package:SummarizedExperiment’:
shift
The following object is masked from ‘package:GenomicRanges’:
shift
The following object is masked from ‘package:IRanges’:
shift
The following objects are masked from ‘package:S4Vectors’:
first, second
Loading required package: Rcpp
Warning message:
“package ‘Rcpp’ was built under R version 4.3.3”
Attaching package: ‘DT’
The following object is masked from ‘package:SeuratObject’:
JS
The following object is masked from ‘package:Seurat’:
JS
Warning message:
“package ‘reshape2’ was built under R version 4.3.3”
Attaching package: ‘reshape2’
The following objects are masked from ‘package:data.table’:
dcast, melt
The following object is masked from ‘package:tidyr’:
smiths
Attaching package: ‘scales’
The following object is masked from ‘package:purrr’:
discard
The following object is masked from ‘package:readr’:
col_factor
# --- 输入参数配置 ---
## rds_path:输入的 Seurat 对象路径
rds_path = "./25020230_nao_CS.rds"
## colName:Seurat 对象 metadata 中样本列名
colName = "Sample"
## sample_name:需要分析的样本名称。
## - 单样本模式:填入单个样本名即可(例如:"sample1")。
## - 多样本整合模式:填入多个样本名,用逗号分隔(例如:"sample1,sample2,sample3")。系统会自动触发多样本整合与批次校正。
sample_name = "25020230_nao_CS_expression"
## meta_path:外部 metadata 文件路径
meta_path = "./meta.tsv"
## npcs:PCA 降维时使用的的主成分数量
npcs = "30"
## lambda:空间权重参数,控制空间特征和表达特征的相对贡献,多个值用逗号分隔
lambda = "0.6,0.8"
## algo:聚类算法,可选 "leiden", "louvain", "kmeans", "mclust"
algo = "leiden"
## resolution:聚类分辨率,多个值用逗号分隔(适用于 leiden/louvain)
resolution = "0.4,0.8"
## kmeans.centers:kmeans 聚类数量,多个值用逗号分隔(适用于 kmeans)
kmeans.centers = "5,10,15,20"
## mclust.G:mclust 聚类数量,多个值用逗号分隔(适用于 mclust)
mclust.G = "5,10,15,20"# 参数解析
# 解析并处理传入的各项分析参数,如空间权重 (`lambda`)、聚类分辨率 (`resolution`) 等。
if(class(lambda) == 'character'){
lambda = as.numeric(strsplit(as.character(lambda),",")[[1]])
}
if(class(resolution) == 'character'){
resolution = as.numeric(strsplit(as.character(resolution),",")[[1]])
}
if(class(kmeans.centers) == 'character'){
kmeans.centers = as.numeric(strsplit(as.character(kmeans.centers),",")[[1]])
}
if(class(mclust.G) == 'character'){
mclust.G = as.numeric(strsplit(as.character(mclust.G),",")[[1]])
}
npcs = as.numeric(npcs)
if (any(resolution == "")) {
resolution = NULL
}else if (any(kmeans.centers == "")) {
kmeans.centers = NULL
}else if (any(mclust.G == "")) {
mclust.G = NULL
}
if(is.null(resolution)){
rm(resolution)
}else if(is.null(kmeans.centers)){
rm(kmeans.centers)
}else if(is.null(mclust.G)){
rm(mclust.G)
}
sample_name = strsplit(sample_name,",")[[1]]💡 Note
多样本坐标处理注意:在进行多样本整合时,由于各个切片原始的物理坐标往往会发生重叠。因此在提取坐标信息时,本代码会根据样本标识对不同切片的坐标点进行自动平移(加上特定的偏移量),从而将它们在同一张虚拟画布上平铺展开,以防计算空间邻域网络时不同切片的细胞发生错误关联。
# --- 数据加载与坐标处理 ---
# 读取 Seurat 对象及元数据,提取细胞空间坐标,并根据样本进行坐标的标准化和平移处理。
obj<-readRDS(rds_path)
meta = read.delim(meta_path)
obj@meta.data = meta
if(colnames(meta)[1]=="barcodes"){
obj@meta.data = column_to_rownames(obj@meta.data,"barcodes")
} else {
obj@meta.data = column_to_rownames(obj@meta.data,"barcode")
}
columns_to_keep <- colnames(obj@meta.data)[!grepl("clust_|HarBsy_", colnames(obj@meta.data))]
obj@meta.data = obj@meta.data[,columns_to_keep]
obj_sample = subset(obj,cells = rownames(obj@meta.data[obj@meta.data[[colName]] %in% sample_name,]))
spatialCoords <- obj_sample@reductions$spatial@cell.embeddings
if(length(sample_name)>1){
spatialCoords = as.data.frame(spatialCoords)
spatialCoords$Sample = obj_sample@meta.data[[colName]]
spatialCoords$sample_id = as.numeric(factor(spatialCoords$Sample, levels = unique(spatialCoords$Sample)))
spatialCoords = spatialCoords[,-which(colnames(spatialCoords) %in% "Sample")]
locs_dt <- data.table(spatialCoords)
colnames(locs_dt) <- c("sdimx", "sdimy", "group")
locs_dt[, sdimx := sdimx - min(sdimx), by = group]
locs_dt$group = as.numeric(locs_dt$group)
global_max <- max(locs_dt$sdimx) * 1.5
locs_dt[, sdimx := sdimx + group * global_max]
locs <- as.matrix(locs_dt[, 1:2])
rownames(locs) <- rownames(obj_sample@meta.data)
} else if(length(sample_name)==1){
colnames(spatialCoords) <- c("sdimx", "sdimy")
}# --- 对象转换 ---
# 将 Seurat 对象转换为 `SpatialExperiment` 对象,以满足 Banksy 算法的输入格式要求。
sce <- Seurat::as.SingleCellExperiment(obj_sample)
sample_id <- as.character(unique(obj_sample$Sample))
sample <- gsub(sample_id,pattern = "_expression",replacement = "")
# 转为SpatialExperiment对象,该对象作为banksy的输入
if(length(sample_name)>1){
spe = SpatialExperiment(
assays = assays(sce),
rowData = rowData(sce),
colData = colData(sce),
spatialCoords = locs
)
} else if(length(sample_name)==1){
spe = SpatialExperiment(
assays = assays(sce),
rowData = rowData(sce),
colData = colData(sce),
sample_id = sample_id,
spatialCoords = spatialCoords,
)
}# --- 数据归一化与高变基因选择 ---
# 根据数据集大小自动选择处理策略:对于小数据集使用全基因进行归一化,对于大数据集提取高变特征基因 (HVGs) 进行后续分析。
if (dim(spe)[2] <= 50000 & dim(spe)[1] <= 40000) {
# 默认使用全部基因,如果使用全部基因,细胞数必需在50000以内,将数据归一化
spe <- computeLibraryFactors(spe)
aname <- "normcounts"
assay(spe, aname) <- normalizeCounts(spe, log = FALSE)
} else {
# 使用高变基因
seu <- as.Seurat(spe, data = NULL)
seu <- FindVariableFeatures(seu, nfeatures = 2000)
# 标准化
scale_factor <- median(colSums(assay(spe, "counts")))
seu <- NormalizeData(seu, scale.factor = scale_factor, normalization.method = "RC")
# 标准化结果加到SpatialExperiment对象
aname <- "normcounts"
assay(spe, aname) <- GetAssayData(seu)
spe <- spe[VariableFeatures(seu),]
}空间特征计算与聚类核心
Banksy 通过 计算邻域特征、降维 与 空间聚类 完成区域识别。该过程包含以下关键步骤:
- 特征计算:利用空间邻域信息增强基因表达特征。
- 降维:使用 PCA 和 UMAP 在增强特征空间内进行降维。
- 批次效应校正:如果存在多样本,利用 Harmony 算法对多个样本间的空间特征进行批次效应校正。
- 聚类:基于降维结果执行空间微环境聚类。
# --- Banksy 特征计算与降维 ---
# 计算基于空间邻域的 Banksy 增强特征矩阵,并执行 PCA 降维以提取主要空间特征。
k_geom <- c(15, 30)
spe <- Banksy::computeBanksy(spe, assay_name = aname, compute_agf = TRUE, k_geom = k_geom)
lambda <- lambda
set.seed(1000)
spe <- Banksy::runBanksyPCA(spe, use_agf = TRUE, lambda = lambda, npcs = npcs)“sparse->dense coercion: allocating vector of size 10.1 GiB”
Computing neighbors...n
Spatial mode is kNN_median
Parameters: k_geom=15
Done
Computing neighbors...n
Spatial mode is kNN_median
Parameters: k_geom=30
Done
Computing harmonic m = 0
Using 15 neighbors
Done
Computing harmonic m = 1
Using 30 neighbors
Centering
Done
Warning message in asMethod(object):
“sparse->dense coercion: allocating vector of size 10.1 GiB”
Warning message in asMethod(object):
“sparse->dense coercion: allocating vector of size 10.1 GiB”
单样本与多样本整合聚类:
本教程同时支持单样本空间聚类和多样本整合空间聚类。系统会根据您在后续配置中输入的样本数量(sample_name 参数)自动判断分析模式:
- 单样本模式:仅对单个组织切片进行空间特征增强和聚类。
- 多样本模式:针对多个组织切片,在空间特征计算后,自动引入 Harmony 算法进行批次效应校正,随后再进行联合降维与聚类,确保在消除技术批次差异的同时保留真实的空间生物学异质性。
# --- 批次效应校正 (仅多样本模式) ---
if(length(sample_name)>1){
for (i in lambda) {
set.seed(1000)
pca_name <- paste0("PCA_M1_lam", i)
print(paste0("computeBanksy Harmony: ", pca_name))
harmony_embedding <- HarmonyMatrix(
data_mat = reducedDim(spe, pca_name),
meta_data = colData(spe),
vars_use = colName,
do_pca = FALSE,
max.iter.harmony = 20,
verbose = FALSE
)
banksy_name <- paste0("HarBsy_lam", i)
reducedDim(spe, banksy_name) <- harmony_embedding
}
}降维与聚类单/多样本差异处理:
- 多样本整合模式:将基于 Harmony 校正后的特征(
dimred = 'HarBsy_lam...')进行 UMAP 降维与聚类。 - 单样本模式:直接使用未校正的原始 Banksy 空间特征进行后续降维与聚类。
# --- 降维与空间聚类 ---
if(length(sample_name)>1){
print("computeBanksy UMAP")
#spe <- Banksy::runBanksyUMAP(spe, use_agf = TRUE, lambda = lambda, npcs = npcs)
for (i in lambda) {
banksy_name <- paste0("HarBsy_lam", i)
spe <- Banksy::runBanksyUMAP(spe, dimred = banksy_name)
}
for (i in lambda) {
banksy_name <- paste0("HarBsy_lam", i)
if(algo == "leiden"|algo == "louvain"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = 'leiden', npcs = npcs, resolution = resolution,dimred = banksy_name)
}else if(algo == "kmeans"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "kmeans",npcs = npcs, kmeans.centers = kmeans.centers,dimred = banksy_name)
}else if(algo == "mclust"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "mclust",npcs = npcs, mclust.G = mclust.G,dimred = banksy_name)
}
}
spe <- Banksy::connectClusters(spe)
} else if(length(sample_name)==1){
spe <- Banksy::runBanksyUMAP(spe, use_agf = TRUE, lambda = lambda, npcs = npcs)
if(algo == "leiden"|algo == "louvain"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = 'leiden', npcs = npcs, resolution = resolution)
}else if(algo == "kmeans"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "kmeans",npcs = npcs, kmeans.centers = kmeans.centers)
}else if(algo == "mclust"){
spe <- Banksy::clusterBanksy(spe, use_agf = TRUE, lambda = lambda, algo = "mclust",npcs = npcs, mclust.G = mclust.G)
}
spe <- Banksy::connectClusters(spe)
}clust_M1_lam0.8_k50_res0.8 --> clust_M1_lam0.6_k50_res0.4
clust_M1_lam0.6_k50_res0.8 --> clust_M1_lam0.8_k50_res0.8
colData(spe) <- cbind(colData(spe), spatialCoords(spe))
data = as.data.frame(colData(spe))
color_palette_24 = c("#E41A1C", "#377EB8", "#4DAF4A", "#984EA3", "#FF7F00",
"#FFFF33", "#A65628", "#F781BF", "#999999", "#66C2A5",
"#FC8D62", "#8DA0CB", "#E78AC3", "#A6D854", "#FFD92F",
"#E5C494", "#B3B3B3", "#6A3D9A", "#FF4500", "#2E8B57",
"#9467BD", "#8C564B", "#E377C2", "#7F7F7F", "#BCBD22",
"#17BECF", "#A6761D", "#1B9E77", "#D95F02", "#7570B3",
"#E7298A", "#66A61E", "#E6AB02", "#A6761D", "#666666",
"#A6CEE3", "#1F78B4", "#B2DF8A", "#33A02C", "#FB9A99",
"#E31A1C", "#FDBF6F", "#FF7F00", "#CAB2D6", "#6A3D9A",
"#FFFF99", "#B15928", "#BDC3C7", "#2C3E50", "#E67E22")
cnames <- grep("^clust", colnames(colData(spe)), value = TRUE)空间聚类可视化
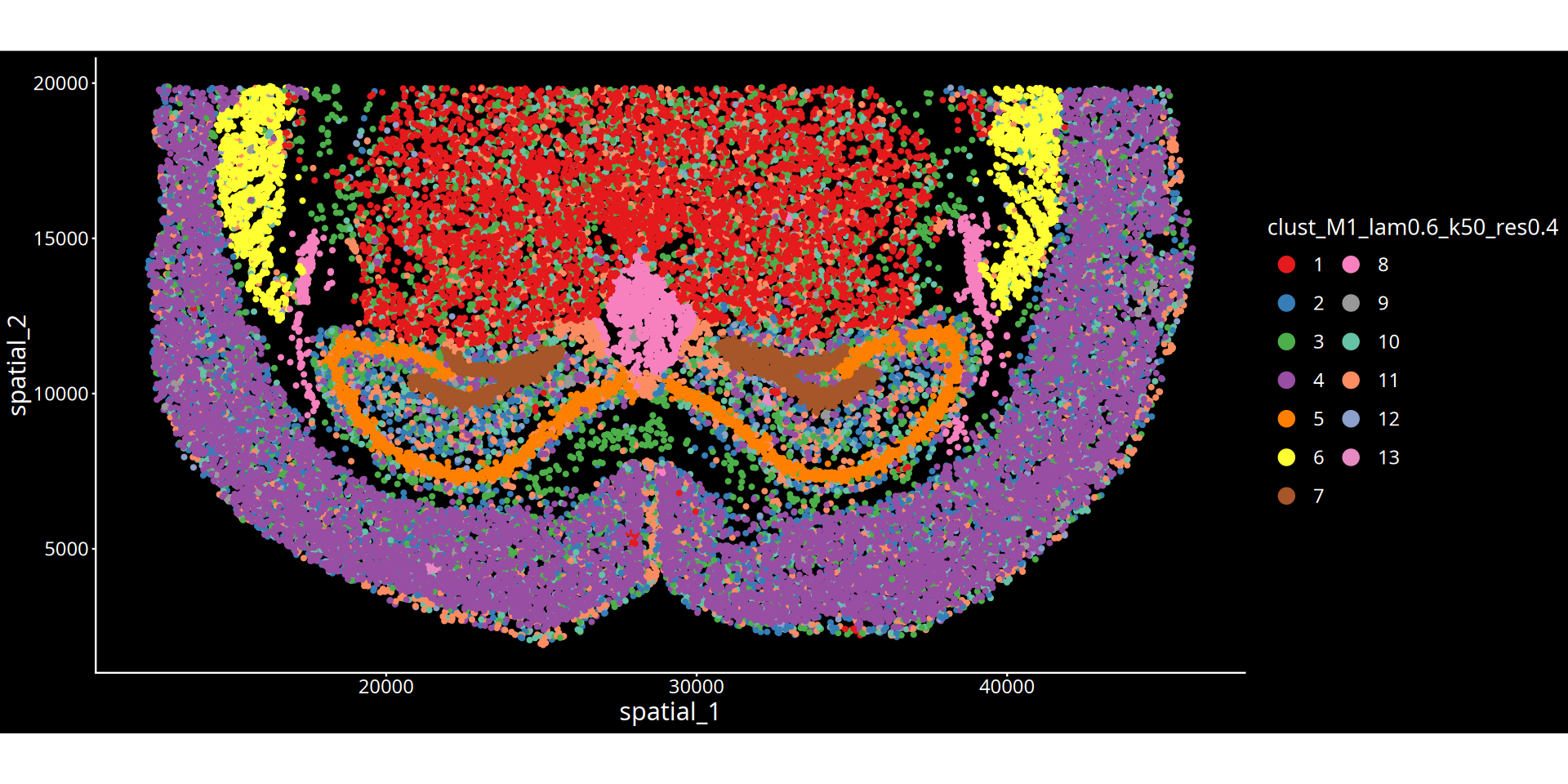
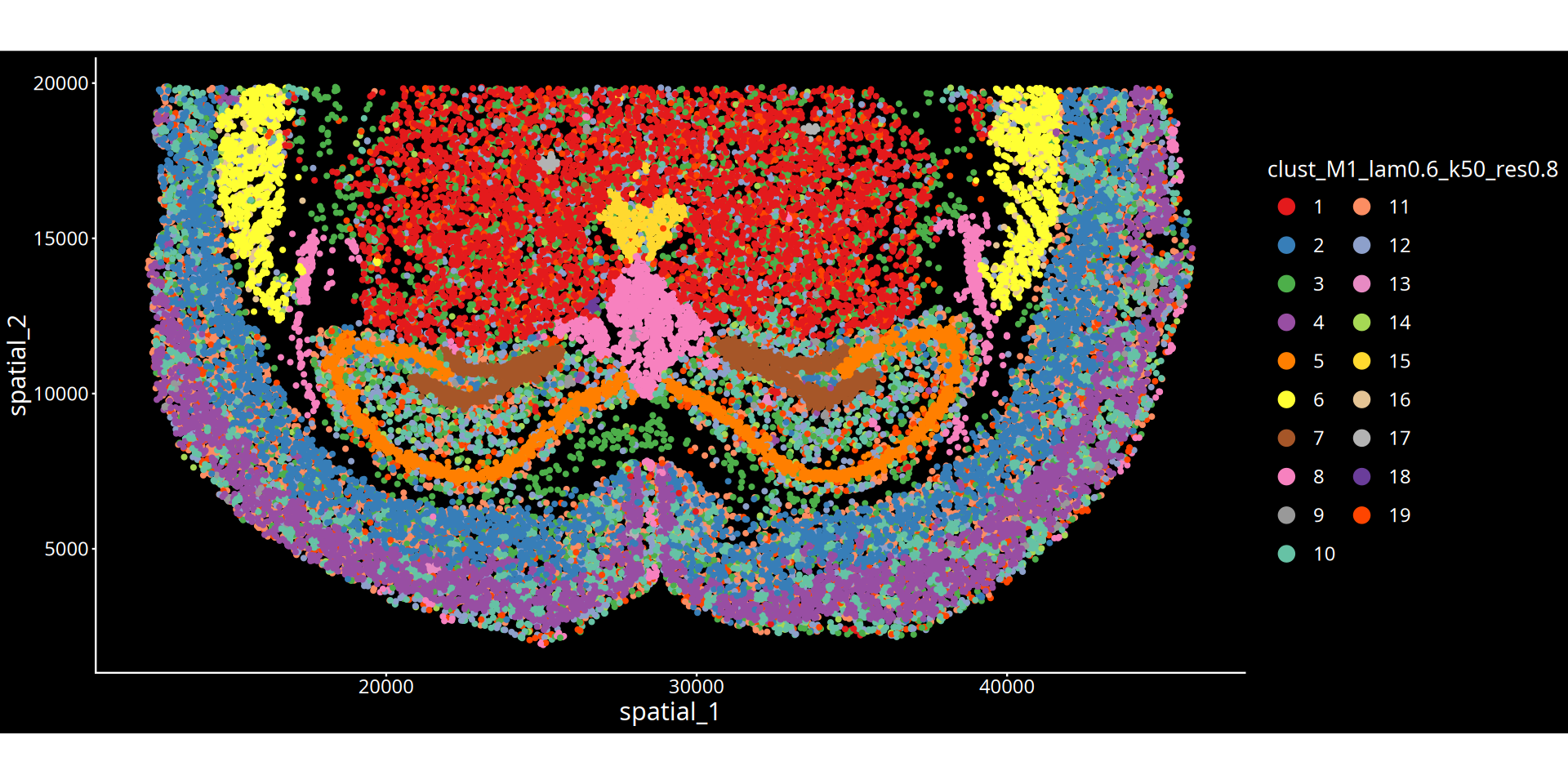
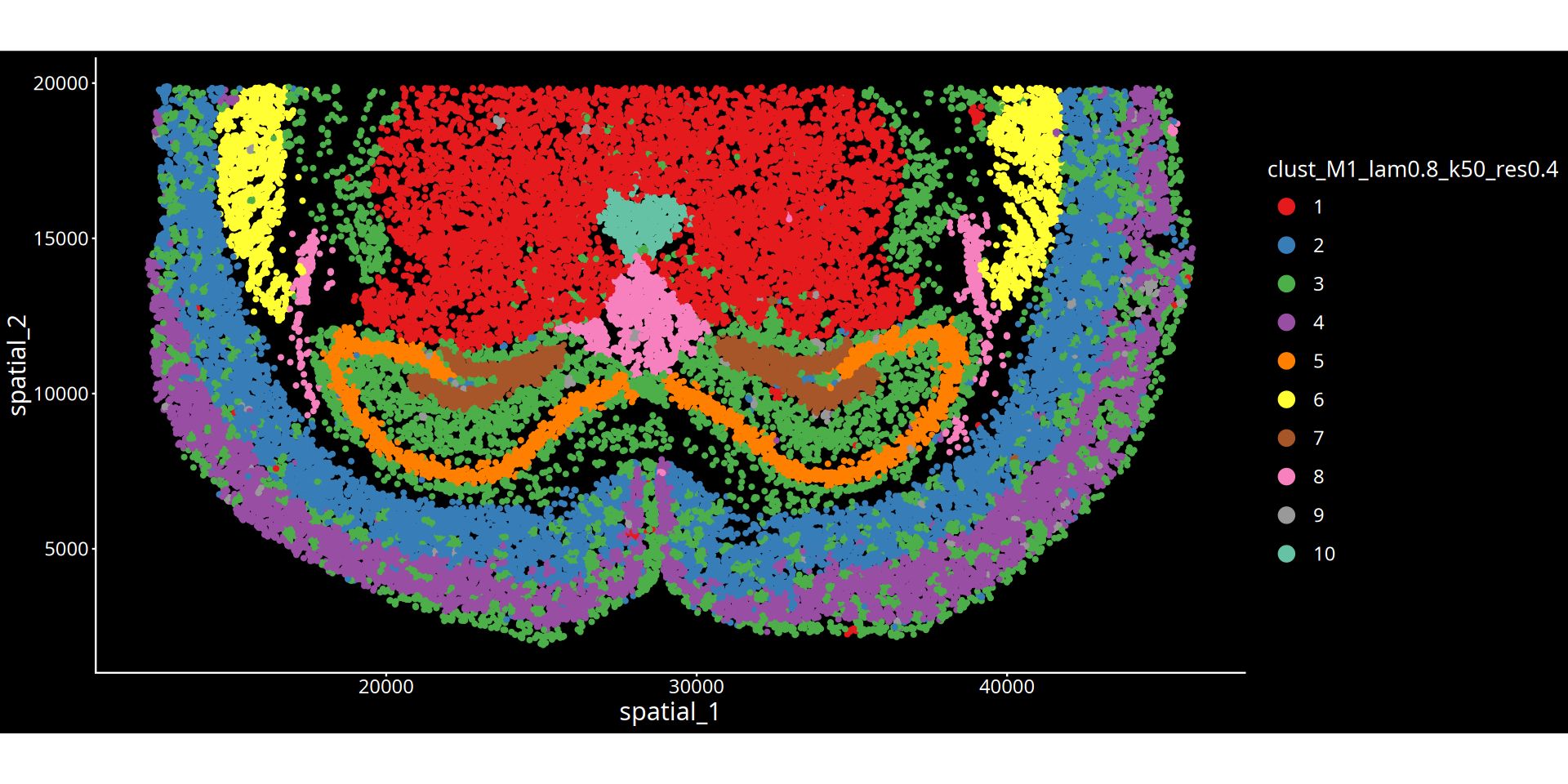
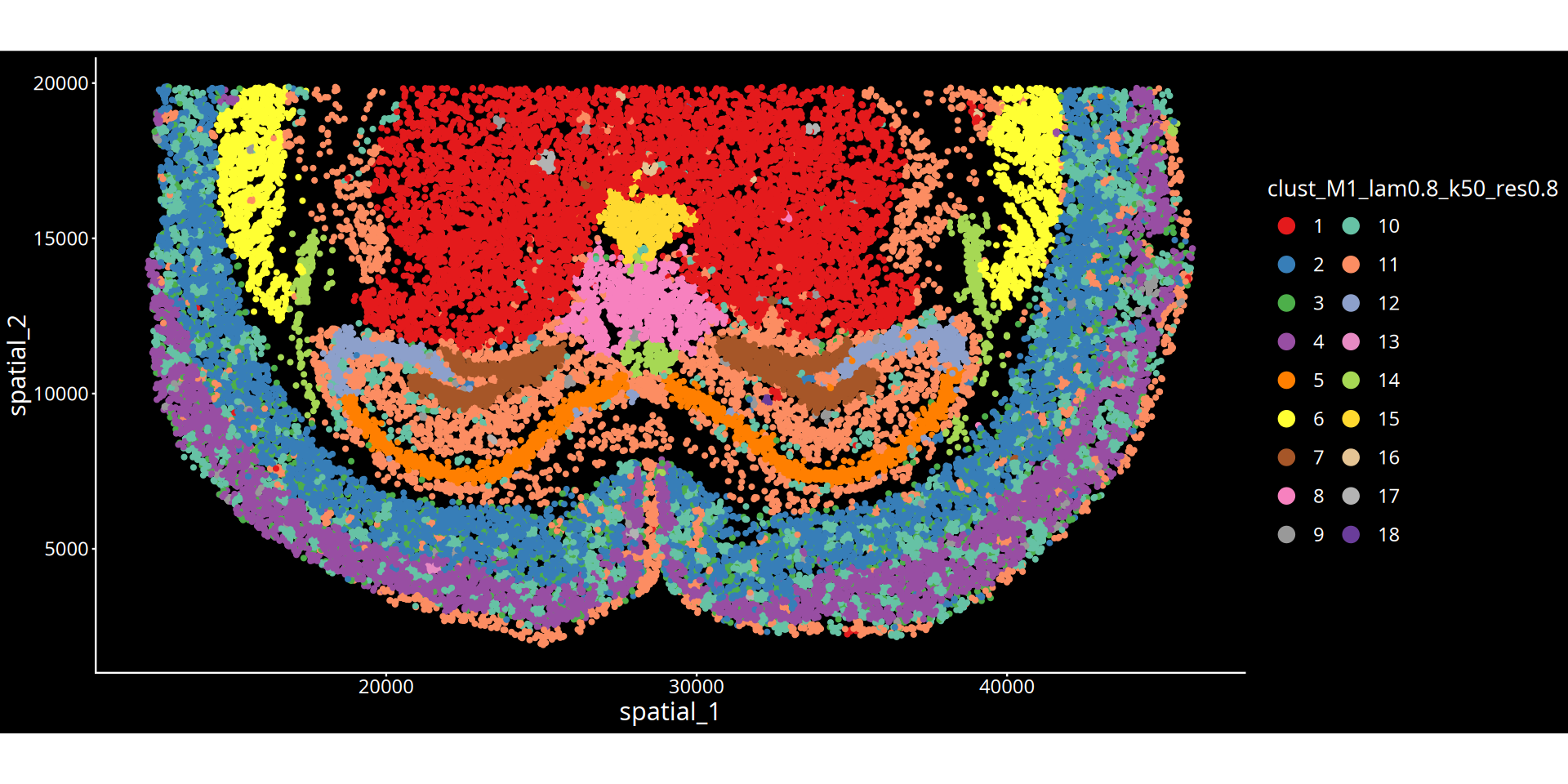
图注说明:下图展示组织切片的空间聚类图。
- 坐标轴:代表组织切片的物理空间坐标,映射细胞在真实组织中的位置。
- 颜色标记:不同的颜色区分了不同的空间聚类簇(Cluster),每一个簇可以理解为具有特定分子特征和空间邻近性的组织微环境(或生态位)。
- 图名参数说明:
- lam:表示空间权重参数(λ),决定了细胞自身表达与周围微环境特征的混合比例。λ 值越大,聚类结果越倾向于考虑空间邻域信息,使得物理位置相近的细胞更容易被聚为一类,形成更平滑连续的空间域;λ 值越小,则越依赖细胞自身的转录组特征。
- 分群参数:根据所选算法不同,
res后的数值表示 leiden/louvain 算法的聚类分辨率;kmeans/mclust后的数值表示预设的分群数量。💡 分析建议与优化: 空间分群的精细程度受空间权重(λ)与分群参数(res/kmeans/mclust)共同控制。若当前分群结果过于细碎,可在后续的云平台交互界面中,结合解剖学结构手动合并相邻或相似的区域。
options(repr.plot.height=8, repr.plot.width=16)
for(i in 1:length(sample_name)){
a = data[data$Sample %in% sample_name[i],]
for(j in 1:length(cnames)){
# 动态计算图例列数
num_clusters <- length(unique(a[[cnames[j]]])) # 获取当前cluster变量的类别数
if(num_clusters <= 10){
ncol_legend <- 1
} else if(num_clusters <= 20){
ncol_legend <- 2
} else if(num_clusters <= 30){
ncol_legend <- 3
} else {
ncol_legend <- 4 # 超过30个时使用4列
}
plots = ggplot(a, aes(sdimx, sdimy))+
geom_point(aes_string(color=cnames[j], fill = cnames[j]), size = 1.2)+
coord_equal()+
theme_classic()+
theme(plot.background = element_rect(fill = "black", color = NA),
panel.background = element_rect(fill = "black", color = NA),
legend.background = element_rect(fill = "black", color = NA),
legend.key = element_rect(fill = "black", color = NA),
text = element_text(color = "white", size = 16),
axis.text = element_text(color = "white", size = 14),
axis.title = element_text(color = "white", size = 18),
axis.line = element_line(color = "white"),
axis.ticks = element_line(color = "white"),
legend.title = element_text(color = "white", size = 16),
legend.text = element_text(color = "white", size = 14),
legend.position = "right",
legend.spacing = unit(0.5, "cm"),
legend.key.size = unit(1, "cm")) +
scale_color_manual(values = color_palette_24)+
xlab("spatial_1")+
ylab("spatial_2")+
guides(color = guide_legend(override.aes = list(size = 5),
ncol = ncol_legend)) # 使用动态计算的列数
print(plots)
}
}
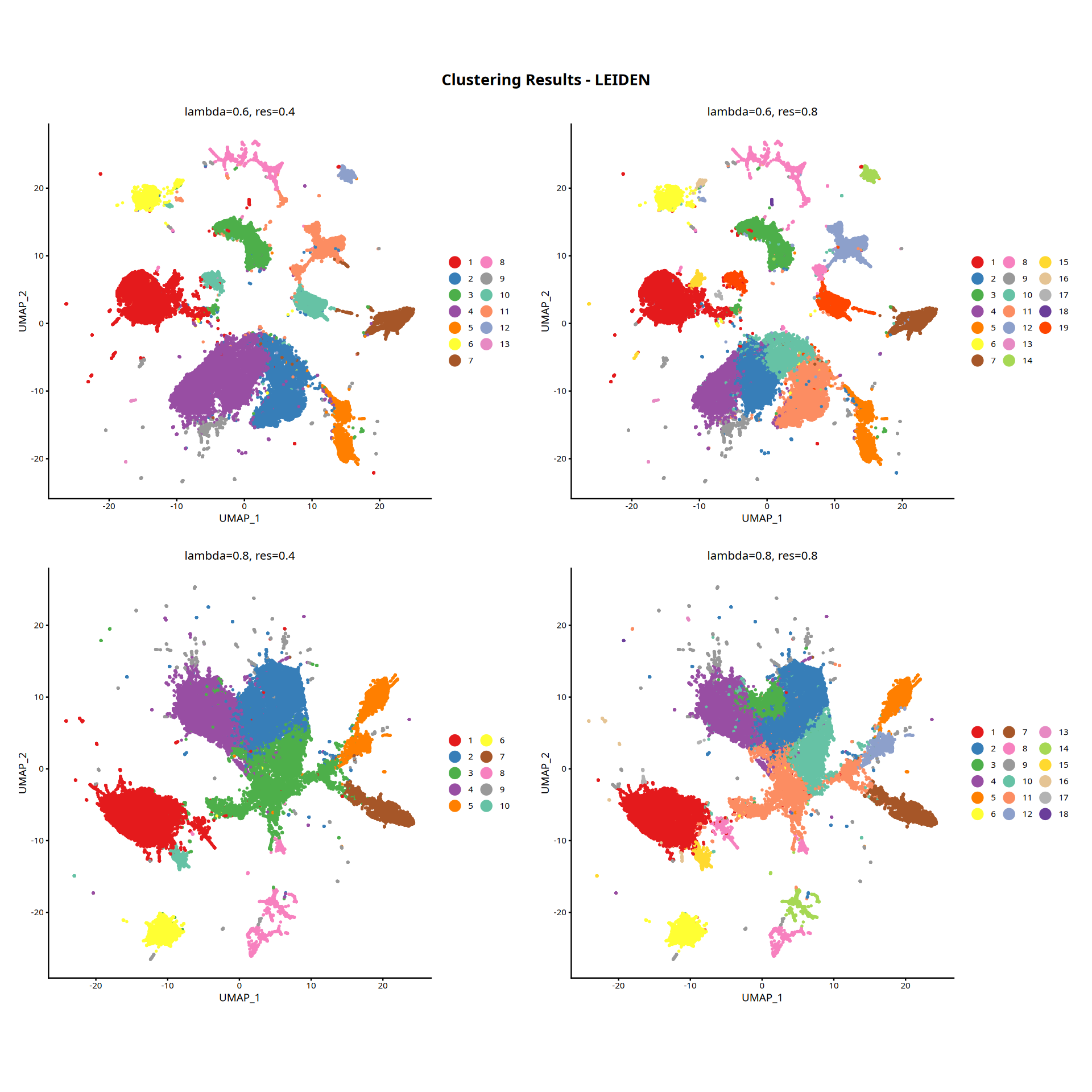
图注说明:下图展示细胞在降维空间中的 UMAP 图。
- 横/纵坐标:UMAP 降维后的二维坐标(UMAP_1 与 UMAP_2),保留高维特征空间的拓扑结构。
- 颜色:空间聚类结果(Cluster)。
# 设置图形参数
options(repr.plot.height=16, repr.plot.width=16)
# 初始化列表存储所有图形
plot_list <- list()
plot_counter <- 1
# 根据样本数量执行不同的处理
if(length(sample_name) > 1){
# 多样本情况
if(algo == "leiden" | algo == "louvain"){
for(i in 1:length(lambda)){
# 获取UMAP降维数据
b = reducedDim(spe, paste0("UMAP_HarBsy_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
paste0("UMAP_HarBsy_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(resolution)){
# 获取聚类列名
cluster_col <- paste0("clust_HarBsy_lam", lambda[i], "_k50_res", resolution[j])
# 计算聚类数量并设置图例列数
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
# 创建图形
plots = ggplot(data, aes_string(x = paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
y = paste0("UMAP_HarBsy_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.x = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", res=", resolution[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend),
fill = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
# 存储图形
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "kmeans"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_HarBsy_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
paste0("UMAP_HarBsy_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(kmeans.centers)){
cluster_col <- paste0("clust_HarBsy_lam", lambda[i], "_k50_kmeans", kmeans.centers[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
y = paste0("UMAP_HarBsy_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.x = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", kmeans=", kmeans.centers[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend),
fill = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "mclust"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_HarBsy_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
paste0("UMAP_HarBsy_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(mclust.G)){
cluster_col <- paste0("clust_HarBsy_lam", lambda[i], "_k50_mclust", mclust.G[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_HarBsy_lam", lambda[i], "_1"),
y = paste0("UMAP_HarBsy_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.x = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", mclust G=", mclust.G[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend),
fill = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
}
} else if(length(sample_name) == 1){
# 单样本情况
if(algo == "leiden" | algo == "louvain"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_M1_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_M1_lam", lambda[i], "_1"),
paste0("UMAP_M1_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(resolution)){
cluster_col <- paste0("clust_M1_lam", lambda[i], "_k50_res", resolution[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_M1_lam", lambda[i], "_1"),
y = paste0("UMAP_M1_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.y = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", res=", resolution[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "kmeans"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_M1_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_M1_lam", lambda[i], "_1"),
paste0("UMAP_M1_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(kmeans.centers)){
cluster_col <- paste0("clust_M1_lam", lambda[i], "_k50_kmeans", kmeans.centers[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_M1_lam", lambda[i], "_1"),
y = paste0("UMAP_M1_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.y = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", kmeans=", kmeans.centers[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
} else if(algo == "mclust"){
for(i in 1:length(lambda)){
b = reducedDim(spe, paste0("UMAP_M1_lam", lambda[i]))
b = as.data.frame(b)
colnames(b) = c(paste0("UMAP_M1_lam", lambda[i], "_1"),
paste0("UMAP_M1_lam", lambda[i], "_2"))
data = cbind(data, b)
for(j in 1:length(mclust.G)){
cluster_col <- paste0("clust_M1_lam", lambda[i], "_k50_mclust", mclust.G[j])
n_clusters <- length(unique(data[[cluster_col]]))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
plots = ggplot(data, aes_string(x = paste0("UMAP_M1_lam", lambda[i], "_1"),
y = paste0("UMAP_M1_lam", lambda[i], "_2"))) +
geom_point(aes_string(color = cluster_col, fill = cluster_col), size = 0.5) +
coord_equal() +
theme_classic() +
theme(legend.position = "right",
legend.box = "vertical",
legend.title = element_blank(),
legend.spacing.y = unit(0.2, "cm"),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm"),
plot.title = element_text(size = 12, hjust = 0.5)) +
scale_color_manual(values = color_palette_24) +
xlab("UMAP_1") +
ylab("UMAP_2") +
ggtitle(paste0("lambda=", lambda[i], ", mclust G=", mclust.G[j])) +
guides(color = guide_legend(override.aes = list(size = 5), ncol = ncol_legend))
plot_list[[plot_counter]] <- plots
plot_counter <- plot_counter + 1
}
}
}
}
# 组合所有图形
# 计算每行显示的图形数量(可以根据需要调整)
ncol_layout <- 2 # 每行显示2个图形
nrow_layout <- ceiling(length(plot_list) / ncol_layout)
# 使用wrap_plots组合图形
combined_plot <- wrap_plots(plot_list, ncol = ncol_layout, byrow = TRUE) +
plot_annotation(
title = paste0("Clustering Results - ", toupper(algo)),
theme = theme(plot.title = element_text(size = 16, hjust = 0.5, face = "bold"))
)
# 显示组合后的图形
print(combined_plot)“annotation$theme is not a valid theme.
Please use \`theme()\` to construct themes.”
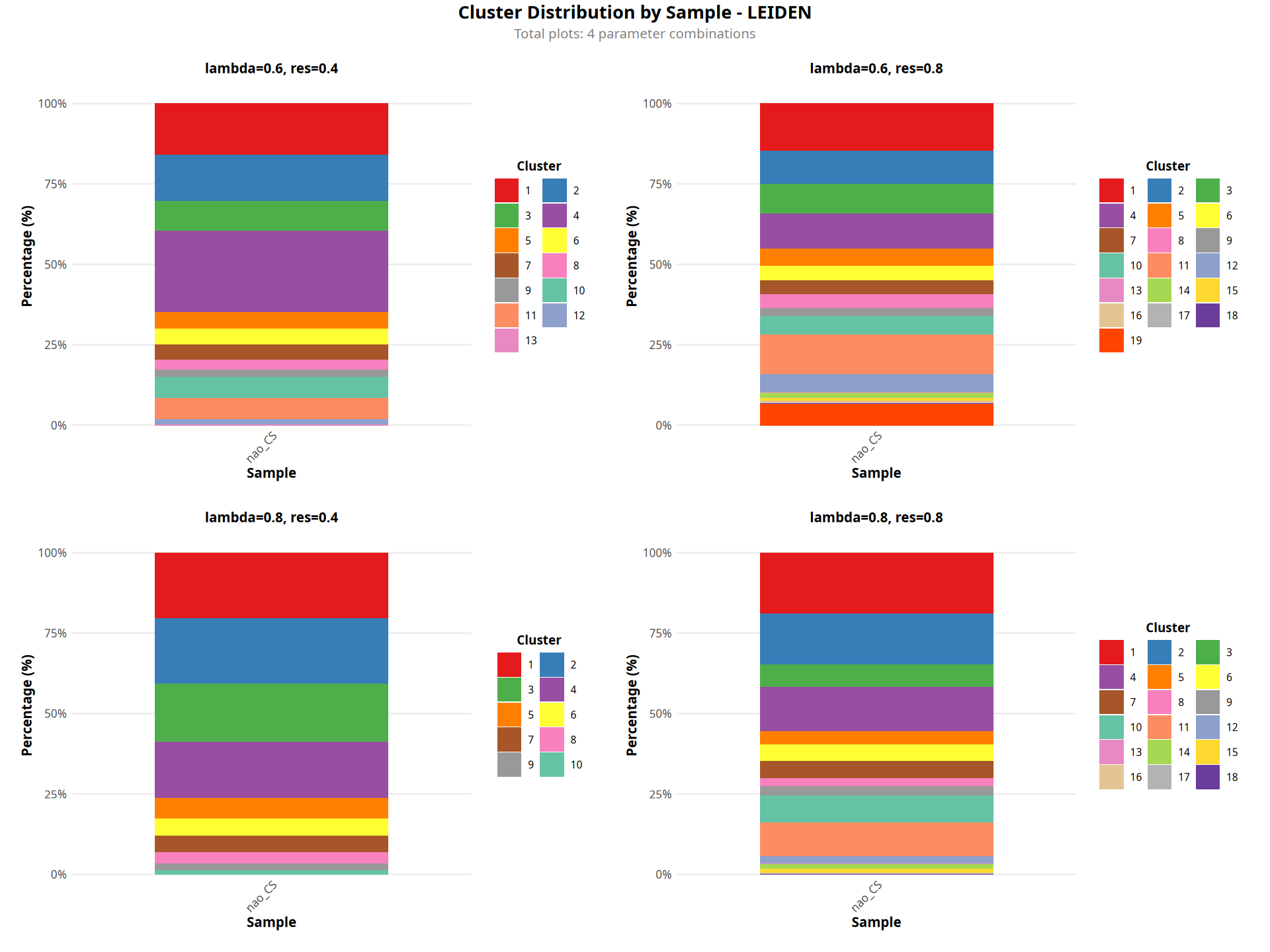
图注说明:下图展示各样本的聚类比例堆叠柱状图。
- 横坐标:样本名称。
- 纵坐标:百分比(Percentage),表示各聚类在样本中的细胞占比。
- 颜色:空间聚类结果(Cluster)。
- 判读:用于比较不同样本之间空间区域组成的差异。
# 加载必要的包
library(ggplot2)
library(reshape2)
library(dplyr)
library(stringr)
library(scales)
library(patchwork)
# 初始化列表存储所有图形
plot_list <- list()
plot_counter <- 1
# 设置图形参数
options(repr.plot.height=12, repr.plot.width=16)
for(i in 1:length(cnames)){
# 提取参数
lam = as.numeric(gsub(".*lam([0-9.]+)_.*", "\\1", cnames[i]))
if(algo == "leiden" | algo == "louvain"){
res = as.numeric(gsub(".*res([0-9.]+).*", "\\1", cnames[i]))
param_label <- paste0("lambda=", lam, ", res=", res)
} else if(algo == "kmeans"){
km = as.numeric(gsub(".*kmeans([0-9]+).*", "\\1", cnames[i]))
param_label <- paste0("lambda=", lam, ", kmeans=", km)
} else if(algo == "mclust"){
g = as.numeric(gsub(".*mclust([0-9]+).*", "\\1", cnames[i]))
param_label <- paste0("lambda=", lam, ", mclust G=", g)
}
# 从table命令创建基础数据
tab <- table(data[[colName]], data[[cnames[i]]])
# 转换为数据框
df <- as.data.frame.matrix(tab)
df$Sample <- rownames(df)
# 转换为长格式
df_long <- melt(df, id.vars = "Sample", variable.name = "Cluster", value.name = "Count")
df_long$Sample = str_replace_all(df_long$Sample, c("^\\d+_" = "", "_expression.*$" = ""))
# 计算百分比并按样本分组
df_long <- df_long %>%
group_by(Sample) %>%
mutate(
Percentage = Count / sum(Count) * 100,
Percentage_label = sprintf("%.1f%%", Percentage)
) %>%
filter(Percentage > 0) %>%
arrange(Sample, desc(Percentage))
# 获取聚类数量并自动设置图例列数
n_clusters <- length(unique(df_long$Cluster))
ncol_legend <- case_when(
n_clusters <= 8 ~ 1,
n_clusters <= 16 ~ 2,
n_clusters <= 24 ~ 3,
n_clusters <= 32 ~ 4,
n_clusters <= 40 ~ 5,
TRUE ~ 6
)
# 绘制图形
p <- ggplot(df_long, aes(x = Sample, y = Percentage, fill = Cluster)) +
geom_bar(stat = "identity", position = "stack", width = 0.7) +
scale_fill_manual(values = color_palette_24[1:n_clusters]) +
labs(
title = param_label,
x = "Sample",
y = "Percentage (%)"
) +
scale_y_continuous(
labels = percent_format(scale = 1),
expand = expansion(mult = c(0, 0.05))
) +
theme_minimal() +
theme(
# X轴标签
axis.text.x = element_text(angle = 45, hjust = 1, size = 10),
axis.title.x = element_text(size = 12, face = "bold"),
# Y轴标签
axis.text.y = element_text(size = 10),
axis.title.y = element_text(size = 12, face = "bold"),
# 标题
plot.title = element_text(size = 12, face = "bold", hjust = 0.5, margin = margin(b = 10)),
# 图例设置
legend.position = "right",
legend.title = element_text(size = 11, face = "bold"),
legend.text = element_text(size = 9),
legend.key.size = unit(0.8, "cm"),
legend.spacing.y = unit(0.2, "cm"),
legend.margin = margin(t = 10, r = 10, b = 10, l = 10),
# 网格线
panel.grid.major.x = element_blank(),
panel.grid.minor = element_blank(),
# 背景和边距
panel.background = element_rect(fill = "white", color = NA),
plot.background = element_rect(fill = "white", color = NA),
plot.margin = unit(c(0.5, 0.5, 0.5, 0.5), "cm")
) +
guides(
fill = guide_legend(
ncol = ncol_legend,
byrow = TRUE,
title = "Cluster",
title.position = "top",
title.hjust = 0.5,
label.position = "right",
label.hjust = 0
)
)
# 存储图形
plot_list[[plot_counter]] <- p
plot_counter <- plot_counter + 1
}
# 组合所有图形
# 计算布局(每行显示2-3个图形)
ncol_layout <- ifelse(length(plot_list) >= 6, 3, 2)
nrow_layout <- ceiling(length(plot_list) / ncol_layout)
# 使用wrap_plots组合图形
combined_plot <- wrap_plots(plot_list, ncol = ncol_layout, byrow = TRUE) +
plot_annotation(
title = paste0("Cluster Distribution by Sample - ", toupper(algo)),
subtitle = paste0("Total plots: ", length(plot_list), " parameter combinations"),
theme = theme(
plot.title = element_text(size = 16, hjust = 0.5, face = "bold"),
plot.subtitle = element_text(size = 12, hjust = 0.5, color = "gray50")
)
)
# 显示组合后的图形
print(combined_plot)“annotation$theme is not a valid theme.
Please use \`theme()\` to construct themes.”