Cell Proportion Difference Analysis: Box Plot of Cell Proportions Between Groups
Time: 8 min
Words: 1.4k words
Updated: 2026-03-02
Reads: 0 times
Group Proportion Plot
Parameters to Modify
The AYxxx after data in Seurat.ob and meta read files can be modified according to your annotation workflow
R
seurat.ob = readRDS("data/AY1734579787477/input.rds")
meta <- read.table("data/AY1734579787477/meta.tsv", header = T, sep = "\t", check.names = F)
seurat.ob <- AddMetaData(seurat.ob, meta)
DefaultAssay(seurat.ob) = "RNA"Verification statistics based on cell sample proportions. Percentage is the proportion of each cell type in the sample. cluster_obj requires all cell types as input.
R
prefix="Myeloid" # Save image name prefix
group_name="Tissue"
groups_com = "AdjacentvsTumor"
cluster_name = "CellAnnotation"
cluster_obj = "Monocyte,Macrophage,DC" # Note the order
sample_name="Sample"
outdir="./"
dosave_fig="F" # Whether to save image. Set to F to view single group results only. Must write F to cancel.
dogroup_test="T" # Whether to add inter-group verification. Supports two groups. Can cancel for more than two groups. Must write F to cancel.Run Code and Display Images
No modification needed unless necessary
R
# Filter data
if (groups_com != ""){
group_list=unlist(strsplit(groups_com,"vs"))
sub = subset(seurat.ob,subset = !!sym(group_name) %in% group_list)
}
if (cluster_obj != ""){
cluster_list = unlist(strsplit(cluster_obj,","))
sub = subset(seurat.ob,subset = !!sym(cluster_name) %in% cluster_list)
}
# Calculate proportion of each cell type in each sample
cellratio = prop.table(table(sub@meta.data[[cluster_name]],sub@meta.data[[sample_name]]),margin = 2)
cellratio <- data.frame(cellratio)
cellper <- dcast(cellratio,Var2~Var1,value.var = "Freq")
rownames(cellper) <- cellper[,1]
#cellper <- cellper[,-1]
# Add group information
samples <- sub@meta.data %>% select(!!sym(sample_name),!!sym(group_name)) %>% distinct()
colnames(samples) <- c("Sample","Group")
rownames(samples) <- samples$Sample
cellper$Sample <- samples[rownames(cellper),"Sample"]
cellper$Group <- samples[rownames(cellper),"Group"]
# Plot
pplist = list()
for (group_ in cluster_list){
cellper_ = cellper %>% select(one_of(c('Sample','Group',group_)))
colnames(cellper_) = c('Sample','Group','percent')
cellper_$percent = as.numeric(cellper_$percent)
cellper_ <- cellper_ %>% group_by(Group) %>% mutate(upper=quantile(percent,0.75),lower=quantile(percent,0.25),mean=mean(percent),median=median(percent))
cellper1 = cellper_ %>% filter(Group %in% group_list)
cellper1$Group = factor(cellper1$Group,levels = group_list)
pp1 = ggplot(cellper1,aes(x=Group,y=percent,fill=Group)) +
geom_boxplot()+theme_bw()+
geom_jitter(shape = 21,aes(fill=Group),width = 0.25,height = 0) +
theme(axis.text = element_text(size = 14),axis.title = element_text(size = 16),legend.text = element_text(size = 18),
legend.title = element_text(size = 20),plot.title = element_text(size = 18,face = 'plain'),panel.grid=element_blank()) +
labs(title = group_,y='Percentage')
if (dogroup_test != "F"){
labely = max(cellper1$percent)
my_comparisons <- strsplit(groups_com, split = "vs")
pp1 = pp1 + stat_compare_means(comparisons = my_comparisons,size = 3,method = "t.test")
}
pplist[[group_]] = pp1
}
if (length(pplist) >1){
adjust_size <- function(n_plots, base_width = 15, base_height = 10) {
ncol <- ceiling(sqrt(n_plots))
nrow <- ceiling(n_plots / ncol)
width <- base_width * ncol / 3 +2
height <- base_height * nrow / 2
list(width = width, height = height)
}
remove_x_axis <- function(p) {
p + theme(axis.title.x = element_blank(),axis.text.x = element_blank(),axis.ticks.x = element_blank(),axis.title.y = element_blank())
}
y_title <- ggplot() +
labs(y = "Percent(%)") +
theme(
plot.margin = margin(0, 0, 0, 10),
axis.title.x = element_blank(),
axis.text.x = element_blank(),
axis.ticks.x = element_blank(),
axis.title.y = element_text(angle = 90, hjust = 0.5),
axis.text.y = element_blank(),
axis.ticks.y = element_blank(),
panel.background = element_blank(),
panel.grid = element_blank(),
axis.line = element_blank() ,axis.title = element_text(size = 20)
)
n_plots <- length(pplist)
ncol <- ceiling(sqrt(n_plots))
nrow <- ceiling(n_plots / ncol)
sizes <- adjust_size(n_plots)
pplist_modified <- lapply(pplist, remove_x_axis)
pplist_no_legend <- lapply(pplist_modified, function(p) p + theme(legend.position="none"))
combined_plot <- plot_grid(plotlist = pplist_no_legend, ncol = ncol, align = 'hv')
final_plot <- plot_grid(y_title,combined_plot, get_legend(pp1), rel_widths = c(0.1,1, 0.2),nrow = 1)
if (dosave_fig != "F") {
ggsave(paste0(outdir,"/",prefix,"_",groups_com,"_group_cellratio.png"),final_plot,width = sizes$width, height = sizes$height,type="cairo-png")
ggsave(paste0(outdir,"/",prefix,"_",groups_com,"_group_cellratio.pdf"),final_plot,width = sizes$width, height = sizes$height)
}
}
options(repr.plot.height=sizes$height, repr.plot.width=sizes$width)
print(final_plot)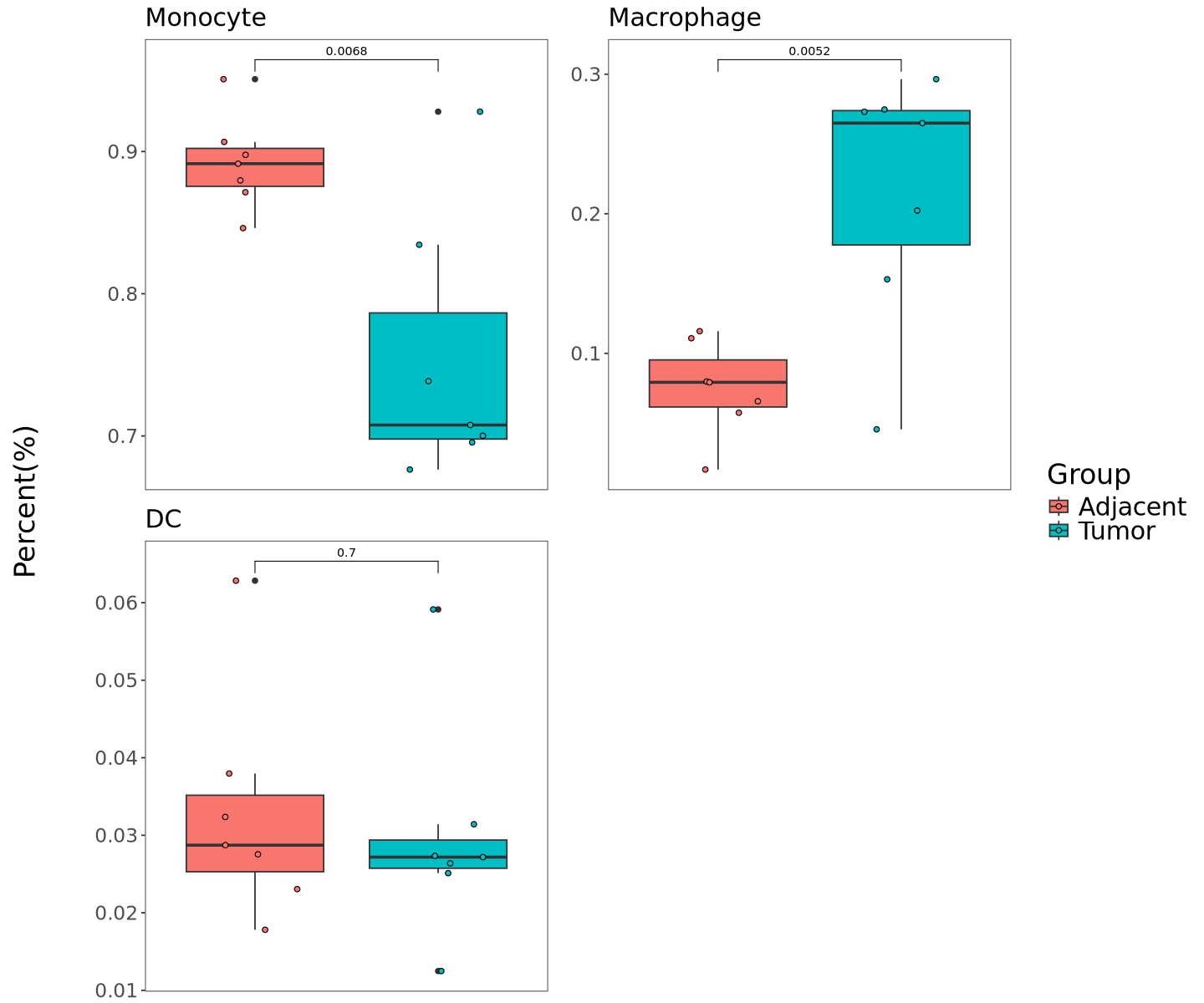
Loop Plotting for Multiple Groups
Define Single Group Plotting Function
R
cellration_barbox <- function(seurat.ob,prefix,group_name,groups_com,cluster_name,cluster_obj,sample_name,outdi,dosave_fig,dogroup_test){
# Filter data
if (groups_com != ""){
group_list=unlist(strsplit(groups_com,"vs"))
sub = subset(seurat.ob,subset = !!sym(group_name) %in% group_list)
}
if (cluster_obj != ""){
cluster_list = unlist(strsplit(cluster_obj,","))
sub = subset(seurat.ob,subset = !!sym(cluster_name) %in% cluster_list)
}
# Calculate proportion of each cell type in each sample
cellratio = prop.table(table(sub@meta.data[[cluster_name]],sub@meta.data[[sample_name]]),margin = 2)
cellratio <- data.frame(cellratio)
cellper <- dcast(cellratio,Var2~Var1,value.var = "Freq")
rownames(cellper) <- cellper[,1]
#cellper <- cellper[,-1]
# Add group information
samples <- sub@meta.data %>% select(!!sym(sample_name),!!sym(group_name)) %>% distinct()
colnames(samples) <- c("Sample","Group")
rownames(samples) <- samples$Sample
cellper$Sample <- samples[rownames(cellper),"Sample"]
cellper$Group <- samples[rownames(cellper),"Group"]
# Plot
pplist = list()
for (group_ in cluster_list){
cellper_ = cellper %>% select(one_of(c('Sample','Group',group_)))
colnames(cellper_) = c('Sample','Group','percent')
cellper_$percent = as.numeric(cellper_$percent)
cellper_ <- cellper_ %>% group_by(Group) %>% mutate(upper=quantile(percent,0.75),lower=quantile(percent,0.25),mean=mean(percent),median=median(percent))
cellper1 = cellper_ %>% filter(Group %in% group_list)
cellper1$Group = factor(cellper1$Group,levels = group_list)
pp1 = ggplot(cellper1,aes(x=Group,y=percent,fill=Group)) +
geom_boxplot()+theme_bw()+
geom_jitter(shape = 21,aes(fill=Group),width = 0.25,height = 0) +
theme(axis.text = element_text(size = 14),axis.title = element_text(size = 16),legend.text = element_text(size = 18),
legend.title = element_text(size = 20),plot.title = element_text(size = 18,face = 'plain'),panel.grid=element_blank()) +
labs(title = group_,y='Percentage')
if (dogroup_test != "F"){
labely = max(cellper1$percent)
my_comparisons <- strsplit(groups_com, split = "vs")
pp1 = pp1 + stat_compare_means(comparisons = my_comparisons,size = 3,method = "t.test")
}
pplist[[group_]] = pp1
}
if (length(pplist) >1){
adjust_size <- function(n_plots, base_width = 15, base_height = 10) {
ncol <- ceiling(sqrt(n_plots))
nrow <- ceiling(n_plots / ncol)
width <- base_width * ncol / 3 +2
height <- base_height * nrow / 2
list(width = width, height = height)
}
remove_x_axis <- function(p) {
p + theme(axis.title.x = element_blank(),axis.text.x = element_blank(),axis.ticks.x = element_blank(),axis.title.y = element_blank())
}
y_title <- ggplot() +
labs(y = "Percent(%)") +
theme(
plot.margin = margin(0, 0, 0, 10),
axis.title.x = element_blank(),
axis.text.x = element_blank(),
axis.ticks.x = element_blank(),
axis.title.y = element_text(angle = 90, hjust = 0.5),
axis.text.y = element_blank(),
axis.ticks.y = element_blank(),
panel.background = element_blank(),
panel.grid = element_blank(),
axis.line = element_blank() ,axis.title = element_text(size = 20)
)
n_plots <- length(pplist)
ncol <- ceiling(sqrt(n_plots))
nrow <- ceiling(n_plots / ncol)
sizes <- adjust_size(n_plots)
pplist_modified <- lapply(pplist, remove_x_axis)
pplist_no_legend <- lapply(pplist_modified, function(p) p + theme(legend.position="none"))
combined_plot <- plot_grid(plotlist = pplist_no_legend, ncol = ncol, align = 'hv')
final_plot <- plot_grid(y_title,combined_plot, get_legend(pp1), rel_widths = c(0.1,1, 0.2),nrow = 1)
if (dosave_fig != "F") {
ggsave(paste0(outdir,"/",prefix,"_",groups_com,"_group_cellratio.png"),final_plot,width = sizes$width, height = sizes$height,type="cairo-png")
ggsave(paste0(outdir,"/",prefix,"_",groups_com,"_group_cellratio.pdf"),final_plot,width = sizes$width, height = sizes$height)
}
}
}Modify Parameters and Save Images
R
prefix="Myeloid" # Save image name prefix
group_name="Tissue"
cluster_name = "CellAnnotation"
cluster_obj = "Monocyte,Macrophage,DC" # Note the order
sample_name="Sample"
outdir="./"
dosave_fig="T" # Whether to save image. Set to F to view single group results only. Must write F to cancel.
dogroup_test="T" # Whether to add inter-group verification. Supports two groups. Can cancel for more than two groups. Must write F to cancel.
groups_coms="AdjacentvsTumor,TumorvsAdjacent"
# No modification needed below
groups_coms_list=unlist(strsplit(groups_coms,","))
for (groups_com in groups_coms_list){
cellration_barbox(seurat.ob,prefix,group_name,groups_com,cluster_name,cluster_obj,sample_name,outdi,dosave_fig,dogroup_test)
}Related images will be stored in the path specified by outdir