COMMOT Spatial Cell Communication: Signal Inference Based on Optimal Transport Algorithm
Background
(1) First, use non-probabilistic mass distribution to control the margins of the transport plan to maintain comparability between species;
(2) Second, enforce spatial distance constraints on CCC to avoid connecting spatially distant parts;
(3) Finally, transport multi-species distributions (ligands) to multi-species distributions (receptors) to explain multi-species interactions.
COMMOT Calculation Results
%%capture
import os
import gc
import ot
import pickle
import anndata
import scanpy as sc
import pandas as pd
import numpy as np
from scipy import sparse
from scipy.stats import spearmanr, pearsonr
from scipy.spatial import distance_matrix
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.colors import Normalize
import commot as ct
import stlearn as st
from PIL import Image
import plotly
import sys
import logging
import re
import warningsTo enable them in other operations, rebuild TensorFlow with the appropriate compiler flags.
2025-09-02 09:50:38.107938: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libcudart.so.11.0'; dlerror: libcudart.so.11.0: cannot open shared object file: No such file or directory
2025-09-02 09:50:38.107975: I tensorflow/compiler/xla/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
2025-09-02 09:51:03.232851: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libnvinfer.so.7'; dlerror: libnvinfer.so.7: cannot open shared object file: No such file or directory
2025-09-02 09:51:03.234823: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libnvinfer_plugin.so.7'; dlerror: libnvinfer_plugin.so.7: cannot open shared object file: No such file or directory
2025-09-02 09:51:03.234839: W tensorflow/compiler/tf2tensorrt/utils/py_utils.cc:38] TF-TRT Warning: Cannot dlopen some TensorRT libraries. If you would like to use Nvidia GPU with TensorRT, please make sure the missing libraries mentioned above are installed properly.
COMMOT Cell Communication Module Input Parameter Introduction
- SAMple_name: Sample name
- meta_path: Path to annotated cell type file, file needs barcode as row name, column name "celltype" as column
- species: Species, related to parameter lr_database_bool below. If lr_database_bool is True, can only be "human" or "mouse"; if False, no need to fill
- lr_database: Fill in COMMOT built-in database name, optional "CellChat" or "CellPhoneDB_v4.0"; if parameter provide_lr_database is True, set name
- signaling_type: If lr_database is CellChat, optional 'Secreted Signaling', 'Cell-Cell Contact', 'ECM-Receptor'; if lr_database is CellPhoneDB_v4.0, optional 'Secreted Signaling', 'Cell-Cell Contact'; if None, use all signals from selected lr_database.
- col_SAM: Sample column name in metadata
- col_celltype: Cell type column name in metadata
sample_name = "N1"
meta_path = "../../data/AY1748480899609/meta.tsv"
species = "human"
lr_database = "CellChat"
signaling_type = None
col_sam="Sample"
col_celltype="CellAnnotation"
def get_cmap_qualitative(cmap_name):
if cmap_name == "Plotly":
cmap = plotly.colors.qualitative.Plotly
elif cmap_name == "Alphabet":
cmap = plotly.colors.qualitative.Alphabet
elif cmap_name == "Light24":
cmap = plotly.colors.qualitative.Light24
elif cmap_name == "Dark24":
cmap = plotly.colors.qualitative.Dark24
# Safe and Vivid are strings of form "rbg(...)"
# Handle this later.
elif cmap_name == "Safe":
cmap = plotly.colors.qualitative.Safe
elif cmap_name == "Vivid":
cmap = plotly.colors.qualitative.Vivid
return cmap- Read SeekSpace data and perform normalization, dimensionality reduction, clustering
adata = sc.read_10x_mtx("filtered_feature_bc_matrix/")
spatial = pd.read_csv('filtered_feature_bc_matrix/cell_locations.tsv',sep="\t",index_col=0)
spatial = spatial.loc[:,("x","y")]
selected_rows = spatial.loc[spatial.index.isin(adata.obs_names)]
selected_rows.columns = ["imagecol","imagerow"]
selected_rows = selected_rows.reindex(adata.obs_names)
selected_rows = selected_rows*0.265385
a = st.create_stlearn(count=adata.to_df(),spatial=selected_rows,library_id=sample_name, scale=1)
a.raw = a
#a.layers["raw_count"] = a.X
# Preprocessing
#st.pp.filter_genes(a,min_cells=3)
st.pp.normalize_total(a)
st.pp.log1p(a)
a_dis500 = a.copy()
# Keep raw data
#a.raw = a
st.pp.scale(a)
st.em.run_pca(a,n_comps=50,random_state=0)
st.pp.neighbors(a,n_neighbors=25,use_rep='X_pca',random_state=0)
st.tl.clustering.louvain(a,random_state=0)
sc.tl.umap(a)Log transformation step is finished in adata.X
Scale step is finished in adata.X
PCA is done! Generated in adata.obsm['X_pca'], adata.uns['pca'] and adata.varm['PCs']
Created k-Nearest-Neighbor graph in adata.uns['neighbors']
Applying Louvain cluster ...n Louvain cluster is done! The labels are stored in adata.obs['louvain']
- Add annotated cell types to adata object
cluster_name = "celltype"
celltype = pd.read_csv(meta_path,index_col=0,sep = "\t")
celltype = celltype.loc[a.obs.index]
a.obs[cluster_name] = celltype[col_celltype]
a.obs[cluster_name] = a.obs[cluster_name].astype('category')
a_dis500 = a_dis500[a.obs.index,:]- Select required receptor-ligand library, filter based on receptor-ligand expression
df_ligrec = ct.pp.ligand_receptor_database(species=species, signaling_type=signaling_type, database=lr_database)
df_ligrec_filtered = ct.pp.filter_lr_database(df_ligrec, a_dis500, min_cell_pct=0.05)
df_ligrec_filtered| 0 | 1 | 2 | 3 | |
|---|---|---|---|---|
| 0 | TGFB1 | TGFBR1_TGFBR2 | TGFb | Secreted Signaling |
| 1 | TGFB2 | TGFBR1_TGFBR2 | TGFb | Secreted Signaling |
| 2 | TGFB3 | TGFBR1_TGFBR2 | TGFb | Secreted Signaling |
| 3 | TGFB1 | ACVR1B_TGFBR2 | TGFb | Secreted Signaling |
| 4 | TGFB2 | ACVR1B_TGFBR2 | TGFb | Secreted Signaling |
| ... | ... | ... | ... | ... |
| 252 | SEMA4G | PLXNB2 | SEMA4 | Cell-Cell Contact |
| 253 | SEMA5A | PLXNA1 | SEMA5 | Cell-Cell Contact |
| 254 | SEMA5A | PLXNA3 | SEMA5 | Cell-Cell Contact |
| 255 | SEMA6A | PLXNA2 | SEMA6 | Cell-Cell Contact |
| 256 | SEMA6A | PLXNA4 | SEMA6 | Cell-Cell Contact |
257 rows × 4 columns
- Infer cell communication for filtered receptor-ligands, distance range 500um.
ct.tl.spatial_communication(a_dis500,
database_name=lr_database, df_ligrec=df_ligrec_filtered, dis_thr=500, heteromeric=True, pathway_sum=True)a_dis500.write("./adata_dis500.h5ad")Number of Signals Sent and Received by Receptor-Ligand Pairs
pts = a_dis500.obsm['spatial']
sender = a_dis500.obsm['commot-' + lr_database + '-sum-sender'][a_dis500.obsm['commot-' + lr_database + '-sum-sender'].columns[1]]
receiver = a_dis500.obsm['commot-' + lr_database + '-sum-receiver'][a_dis500.obsm['commot-' + lr_database + '-sum-receiver'].columns[1]]
fig, ax = plt.subplots(1,2, figsize=(20,8))
ax[0].scatter(pts[:,0], pts[:,1], c=sender, s=5, cmap='Blues')
ax[0].set_title(a_dis500.obsm['commot-' + lr_database + '-sum-sender'].columns[1]+'_Sender')
norm_sender = Normalize(vmin=min(sender), vmax=max(sender))
plt.colorbar(cm.ScalarMappable(norm=norm_sender, cmap='Blues'), ax=ax[0])
ax[0].set_aspect("equal")
ax[1].scatter(pts[:,0], pts[:,1], c=receiver, s=5, cmap='Reds')
ax[1].set_title(a_dis500.obsm['commot-' + lr_database + '-sum-receiver'].columns[1]+'_Receiver')
norm_receiver = Normalize(vmin=min(receiver), vmax=max(receiver))
plt.colorbar(cm.ScalarMappable(norm=norm_receiver, cmap='Reds'), ax=ax[1])
ax[1].set_aspect("equal")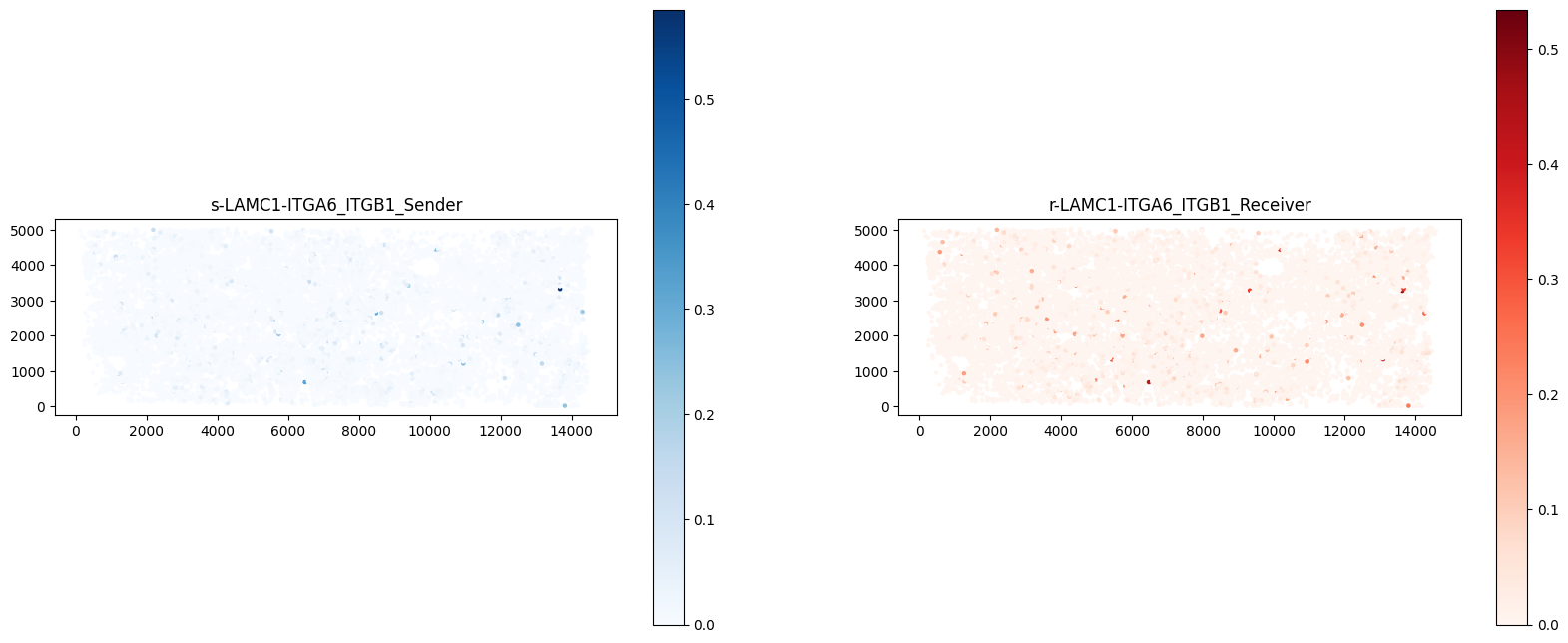
Receptor-ligand co-expression. Left plot represents ligand expression; right plot represents receptor expression; darker color means higher expression. If receptor and ligand are expressed in the same region, it is inferred that interaction occurs between cells expressing this receptor-ligand pair. (Shown here for one receptor-ligand pair)#ct.tl.communication_direction(a_dis500, database_name=lr_database, pathway_name='PSAP', k=5)
#ct.pl.plot_cell_communication(a_dis500, database_name=lr_database, pathway_name='PSAP', plot_method='grid', background_legend=True,
# scale=0.00003, ndsize=8, grid_density=0.4, summary='sender', background='image', clustering=cluster_name, cmap='Alphabet',
# normalize_v = True, normalize_v_quantile=0.995)Cell Communication Signal Flow Visualization
COMMOT considers cell communication as a cell interacting with neighboring cells, which interact with the next neighboring cells, forming a "signal flow" sequentially. The figure shows the signal flow direction of the pathway.
#print("total_signature_stream")
ct.tl.communication_direction(a_dis500, database_name=lr_database, k=5)
ct.pl.plot_cell_communication(a_dis500, database_name=lr_database, lr_pair=('total','total'), plot_method='stream', background_legend=True,
scale=0.00003, ndsize=8, grid_density=0.4, summary='sender', background='summary', clustering=cluster_name, cmap='Reds',
normalize_v = True, normalize_v_quantile=0.995)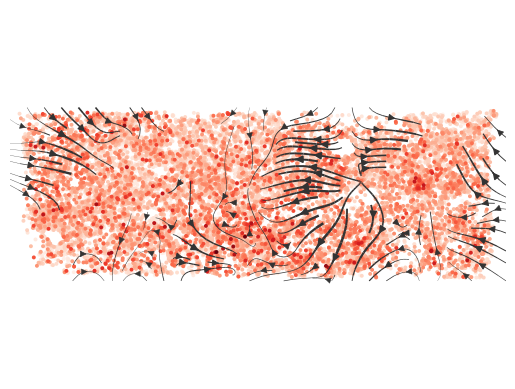
Directionality of communication between each cell. Arrows represent communication direction, background color represents cell signal sending intensity.- Calculate communication intensity between different cell types
a_dis500.obs[cluster_name] = a.obs[cluster_name]
# Calculate results for each receptor-ligand pair
for i,j in enumerate(df_ligrec_filtered.values):
ct.tl.cluster_communication(a_dis500, database_name=lr_database, clustering=cluster_name,n_permutations=100,lr_pair=(j[0],j[1]))
ct.tl.cluster_communication(a_dis500, database_name=lr_database, clustering=cluster_name,n_permutations=100)Cell Communication Network
a_dis500obs: 'imagecol', 'imagerow', 'celltype'
uns: 'spatial', 'log1p', 'commot-CellChat-info', 'commot_cluster-celltype-CellChat-TGFB1-TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB2-TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB3-TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB1-ACVR1B_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB2-ACVR1B_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB3-ACVR1B_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB1-ACVR1_TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB2-ACVR1_TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB3-ACVR1_TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-WNT5B-FZD6', 'commot_cluster-celltype-CellChat-NRG1-ERBB3', 'commot_cluster-celltype-CellChat-NRG1-ERBB2_ERBB3', 'commot_cluster-celltype-CellChat-NRG1-ERBB4', 'commot_cluster-celltype-CellChat-NRG1-ERBB2_ERBB4', 'commot_cluster-celltype-CellChat-NRG2-ERBB3', 'commot_cluster-celltype-CellChat-NRG2-ERBB2_ERBB3', 'commot_cluster-celltype-CellChat-NRG2-ERBB4', 'commot_cluster-celltype-CellChat-NRG2-ERBB2_ERBB4', 'commot_cluster-celltype-CellChat-PDGFA-PDGFRA', 'commot_cluster-celltype-CellChat-PDGFA-PDGFRB', 'commot_cluster-celltype-CellChat-PDGFC-PDGFRA', 'commot_cluster-celltype-CellChat-IGF1-IGF1R', 'commot_cluster-celltype-CellChat-IGF1-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-MIF-CD74_CXCR4', 'commot_cluster-celltype-CellChat-MIF-CD74_CD44', 'commot_cluster-celltype-CellChat-CSF1-CSF1R', 'commot_cluster-celltype-CellChat-NAMPT-INSR', 'commot_cluster-celltype-CellChat-NAMPT-ITGA5_ITGB1', 'commot_cluster-celltype-CellChat-MDK-SDC2', 'commot_cluster-celltype-CellChat-MDK-SDC4', 'commot_cluster-celltype-CellChat-MDK-ITGA4_ITGB1', 'commot_cluster-celltype-CellChat-MDK-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-MDK-LRP1', 'commot_cluster-celltype-CellChat-MDK-NCL', 'commot_cluster-celltype-CellChat-PTN-SDC2', 'commot_cluster-celltype-CellChat-PTN-SDC3', 'commot_cluster-celltype-CellChat-PTN-SDC4', 'commot_cluster-celltype-CellChat-PTN-NCL', 'commot_cluster-celltype-CellChat-C3-ITGAX_ITGB2', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA1', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA2', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA3', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA4', 'commot_cluster-celltype-CellChat-SEMA3C-PLXND1', 'commot_cluster-celltype-CellChat-GAS6-AXL', 'commot_cluster-celltype-CellChat-GAS6-MERTK', 'commot_cluster-celltype-CellChat-GRN-SORT1', 'commot_cluster-celltype-CellChat-LGALS9-PTPRC', 'commot_cluster-celltype-CellChat-LGALS9-CD44', 'commot_cluster-celltype-CellChat-COL1A1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL1A1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGA4_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGA5_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-THBS1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-THBS3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL1A1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-TNC-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-TNXB-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL1A1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGAV_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA3-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA4-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA5-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMB1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMB2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMC1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL1A1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL1A2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A3-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A4-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A5-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A6-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL6A1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL6A2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL6A3-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA2-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMA3-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMA4-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMA5-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMB1-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMB2-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMC1-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-FN1-CD44', 'commot_cluster-celltype-CellChat-COL1A1-CD44', 'commot_cluster-celltype-CellChat-COL1A2-CD44', 'commot_cluster-celltype-CellChat-COL4A1-CD44', 'commot_cluster-celltype-CellChat-COL4A2-CD44', 'commot_cluster-celltype-CellChat-COL4A3-CD44', 'commot_cluster-celltype-CellChat-COL4A4-CD44', 'commot_cluster-celltype-CellChat-COL4A5-CD44', 'commot_cluster-celltype-CellChat-COL4A6-CD44', 'commot_cluster-celltype-CellChat-COL6A1-CD44', 'commot_cluster-celltype-CellChat-COL6A2-CD44', 'commot_cluster-celltype-CellChat-COL6A3-CD44', 'commot_cluster-celltype-CellChat-LAMA2-CD44', 'commot_cluster-celltype-CellChat-LAMA3-CD44', 'commot_cluster-celltype-CellChat-LAMA4-CD44', 'commot_cluster-celltype-CellChat-LAMA5-CD44', 'commot_cluster-celltype-CellChat-LAMB1-CD44', 'commot_cluster-celltype-CellChat-LAMB2-CD44', 'commot_cluster-celltype-CellChat-LAMC1-CD44', 'commot_cluster-celltype-CellChat-COL1A1-SDC4', 'commot_cluster-celltype-CellChat-COL1A2-SDC4', 'commot_cluster-celltype-CellChat-COL4A1-SDC4', 'commot_cluster-celltype-CellChat-COL4A2-SDC4', 'commot_cluster-celltype-CellChat-COL4A3-SDC4', 'commot_cluster-celltype-CellChat-COL4A4-SDC4', 'commot_cluster-celltype-CellChat-COL4A5-SDC4', 'commot_cluster-celltype-CellChat-COL4A6-SDC4', 'commot_cluster-celltype-CellChat-COL6A1-SDC4', 'commot_cluster-celltype-CellChat-COL6A2-SDC4', 'commot_cluster-celltype-CellChat-COL6A3-SDC4', 'commot_cluster-celltype-CellChat-FN1-SDC4', 'commot_cluster-celltype-CellChat-TNC-SDC4', 'commot_cluster-celltype-CellChat-TNXB-SDC4', 'commot_cluster-celltype-CellChat-THBS1-SDC4', 'commot_cluster-celltype-CellChat-THBS3-SDC4', 'commot_cluster-celltype-CellChat-THBS1-CD47', 'commot_cluster-celltype-CellChat-THBS3-CD47', 'commot_cluster-celltype-CellChat-AGRN-DAG1', 'commot_cluster-celltype-CellChat-HSPG2-DAG1', 'commot_cluster-celltype-CellChat-LAMA2-DAG1', 'commot_cluster-celltype-CellChat-LAMA3-DAG1', 'commot_cluster-celltype-CellChat-LAMA4-DAG1', 'commot_cluster-celltype-CellChat-LAMA5-DAG1', 'commot_cluster-celltype-CellChat-LAMB1-DAG1', 'commot_cluster-celltype-CellChat-LAMB2-DAG1', 'commot_cluster-celltype-CellChat-LAMC1-DAG1', 'commot_cluster-celltype-CellChat-APP-CD74', 'commot_cluster-celltype-CellChat-CADM1-CADM1', 'commot_cluster-celltype-CellChat-PTPRC-MRC1', 'commot_cluster-celltype-CellChat-CD46-JAG1', 'commot_cluster-celltype-CellChat-CD96-NECTIN1', 'commot_cluster-celltype-CellChat-CD96-PVR', 'commot_cluster-celltype-CellChat-CD99-CD99', 'commot_cluster-celltype-CellChat-CD99-CD99L2', 'commot_cluster-celltype-CellChat-CDH1-CDH1', 'commot_cluster-celltype-CellChat-CDH1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-EFNA1-EPHA3', 'commot_cluster-celltype-CellChat-EFNA1-EPHA4', 'commot_cluster-celltype-CellChat-EFNA1-EPHA7', 'commot_cluster-celltype-CellChat-EFNA5-EPHA3', 'commot_cluster-celltype-CellChat-EFNA5-EPHA4', 'commot_cluster-celltype-CellChat-EFNA5-EPHA7', 'commot_cluster-celltype-CellChat-EFNB2-EPHA4', 'commot_cluster-celltype-CellChat-EFNB2-EPHB4', 'commot_cluster-celltype-CellChat-F11R-ITGAL_ITGB2', 'commot_cluster-celltype-CellChat-F11R-F11R', 'commot_cluster-celltype-CellChat-F11R-JAM3', 'commot_cluster-celltype-CellChat-JAM3-F11R', 'commot_cluster-celltype-CellChat-JAM3-JAM3', 'commot_cluster-celltype-CellChat-MPZL1-MPZL1', 'commot_cluster-celltype-CellChat-NECTIN1-CD96', 'commot_cluster-celltype-CellChat-NEGR1-NEGR1', 'commot_cluster-celltype-CellChat-DLL1-NOTCH1', 'commot_cluster-celltype-CellChat-DLL1-NOTCH3', 'commot_cluster-celltype-CellChat-JAG1-NOTCH1', 'commot_cluster-celltype-CellChat-JAG1-NOTCH2', 'commot_cluster-celltype-CellChat-JAG1-NOTCH3', 'commot_cluster-celltype-CellChat-DLL1-NOTCH2', 'commot_cluster-celltype-CellChat-NRXN3-NLGN1', 'commot_cluster-celltype-CellChat-NRXN3-NLGN2', 'commot_cluster-celltype-CellChat-PECAM1-PECAM1', 'commot_cluster-celltype-CellChat-PTPRM-PTPRM', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA1', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA2', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA3', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA4', 'commot_cluster-celltype-CellChat-SEMA4A-PLXNB1', 'commot_cluster-celltype-CellChat-SEMA4A-PLXNB2', 'commot_cluster-celltype-CellChat-SEMA4D-PLXNB1', 'commot_cluster-celltype-CellChat-SEMA4D-PLXNB2', 'commot_cluster-celltype-CellChat-SEMA4G-PLXNB2', 'commot_cluster-celltype-CellChat-SEMA5A-PLXNA1', 'commot_cluster-celltype-CellChat-SEMA5A-PLXNA3', 'commot_cluster-celltype-CellChat-SEMA6A-PLXNA2', 'commot_cluster-celltype-CellChat-SEMA6A-PLXNA4', 'commot_cluster-celltype-CellChat-total-total'
obsm: 'spatial', 'commot-CellChat-sum-sender', 'commot-CellChat-sum-receiver', 'commot_sender_vf-CellChat-total-total', 'commot_receiver_vf-CellChat-total-total'
obsp: 'commot-CellChat-LAMC1-ITGA6_ITGB4', 'commot-CellChat-LAMC1-ITGA6_ITGB1', 'commot-CellChat-LAMC1-ITGA2_ITGB1', 'commot-CellChat-LAMC1-ITGA3_ITGB1', 'commot-CellChat-LAMC1-ITGA9_ITGB1', 'commot-CellChat-LAMC1-DAG1', 'commot-CellChat-LAMC1-CD44', 'commot-CellChat-LAMC1-ITGAV_ITGB8', 'commot-CellChat-LAMC1-ITGA1_ITGB1', 'commot-CellChat-LAMB2-ITGA6_ITGB4', 'commot-CellChat-LAMB2-ITGA6_ITGB1', 'commot-CellChat-LAMB2-ITGA2_ITGB1', 'commot-CellChat-LAMB2-ITGA3_ITGB1', 'commot-CellChat-LAMB2-ITGA9_ITGB1', 'commot-CellChat-LAMB2-DAG1', 'commot-CellChat-LAMB2-CD44', 'commot-CellChat-LAMB2-ITGAV_ITGB8', 'commot-CellChat-LAMB2-ITGA1_ITGB1', 'commot-CellChat-NAMPT-INSR', 'commot-CellChat-NAMPT-ITGA5_ITGB1', 'commot-CellChat-COL6A1-SDC4', 'commot-CellChat-COL6A1-ITGA2_ITGB1', 'commot-CellChat-COL6A1-ITGA3_ITGB1', 'commot-CellChat-COL6A1-ITGA9_ITGB1', 'commot-CellChat-COL6A1-CD44', 'commot-CellChat-COL6A1-ITGAV_ITGB8', 'commot-CellChat-COL6A1-ITGA1_ITGB1', 'commot-CellChat-THBS1-CD47', 'commot-CellChat-THBS1-SDC4', 'commot-CellChat-THBS1-ITGA3_ITGB1', 'commot-CellChat-JAM3-JAM3', 'commot-CellChat-JAM3-F11R', 'commot-CellChat-SEMA4G-PLXNB2', 'commot-CellChat-NRXN3-NLGN2', 'commot-CellChat-NRXN3-NLGN1', 'commot-CellChat-COL4A4-SDC4', 'commot-CellChat-COL4A4-ITGA2_ITGB1', 'commot-CellChat-COL4A4-ITGA3_ITGB1', 'commot-CellChat-COL4A4-ITGA9_ITGB1', 'commot-CellChat-COL4A4-CD44', 'commot-CellChat-COL4A4-ITGAV_ITGB8', 'commot-CellChat-COL4A4-ITGA1_ITGB1', 'commot-CellChat-PTPRC-MRC1', 'commot-CellChat-CD99-CD99L2', 'commot-CellChat-CD99-CD99', 'commot-CellChat-TGFB1-ACVR1_TGFBR1_TGFBR2', 'commot-CellChat-TGFB1-TGFBR1_TGFBR2', 'commot-CellChat-TGFB1-ACVR1B_TGFBR2', 'commot-CellChat-COL4A2-SDC4', 'commot-CellChat-COL4A2-ITGA2_ITGB1', 'commot-CellChat-COL4A2-ITGA3_ITGB1', 'commot-CellChat-COL4A2-ITGA9_ITGB1', 'commot-CellChat-COL4A2-CD44', 'commot-CellChat-COL4A2-ITGAV_ITGB8', 'commot-CellChat-COL4A2-ITGA1_ITGB1', 'commot-CellChat-TGFB3-ACVR1_TGFBR1_TGFBR2', 'commot-CellChat-TGFB3-TGFBR1_TGFBR2', 'commot-CellChat-TGFB3-ACVR1B_TGFBR2', 'commot-CellChat-LAMA3-ITGA6_ITGB4', 'commot-CellChat-LAMA3-ITGA6_ITGB1', 'commot-CellChat-LAMA3-ITGA2_ITGB1', 'commot-CellChat-LAMA3-ITGA3_ITGB1', 'commot-CellChat-LAMA3-ITGA9_ITGB1', 'commot-CellChat-LAMA3-DAG1', 'commot-CellChat-LAMA3-CD44', 'commot-CellChat-LAMA3-ITGAV_ITGB8', 'commot-CellChat-LAMA3-ITGA1_ITGB1', 'commot-CellChat-CDH1-ITGA2_ITGB1', 'commot-CellChat-CDH1-CDH1', 'commot-CellChat-LAMA2-ITGA6_ITGB4', 'commot-CellChat-LAMA2-ITGA6_ITGB1', 'commot-CellChat-LAMA2-ITGA2_ITGB1', 'commot-CellChat-LAMA2-ITGA3_ITGB1', 'commot-CellChat-LAMA2-ITGA9_ITGB1', 'commot-CellChat-LAMA2-DAG1', 'commot-CellChat-LAMA2-CD44', 'commot-CellChat-LAMA2-ITGAV_ITGB8', 'commot-CellChat-LAMA2-ITGA1_ITGB1', 'commot-CellChat-EFNA1-EPHA4', 'commot-CellChat-EFNA1-EPHA3', 'commot-CellChat-EFNA1-EPHA7', 'commot-CellChat-CD46-JAG1', 'commot-CellChat-AGRN-DAG1', 'commot-CellChat-C3-ITGAX_ITGB2', 'commot-CellChat-NECTIN1-CD96', 'commot-CellChat-DLL1-NOTCH2', 'commot-CellChat-DLL1-NOTCH1', 'commot-CellChat-DLL1-NOTCH3', 'commot-CellChat-SEMA4D-PLXNB1', 'commot-CellChat-SEMA4D-PLXNB2', 'commot-CellChat-SEMA6A-PLXNA2', 'commot-CellChat-SEMA6A-PLXNA4', 'commot-CellChat-COL6A2-SDC4', 'commot-CellChat-COL6A2-ITGA2_ITGB1', 'commot-CellChat-COL6A2-ITGA3_ITGB1', 'commot-CellChat-COL6A2-ITGA9_ITGB1', 'commot-CellChat-COL6A2-CD44', 'commot-CellChat-COL6A2-ITGAV_ITGB8', 'commot-CellChat-COL6A2-ITGA1_ITGB1', 'commot-CellChat-FN1-ITGA4_ITGB1', 'commot-CellChat-FN1-SDC4', 'commot-CellChat-FN1-ITGA3_ITGB1', 'commot-CellChat-FN1-CD44', 'commot-CellChat-FN1-ITGAV_ITGB8', 'commot-CellChat-FN1-ITGA5_ITGB1', 'commot-CellChat-FN1-ITGAV_ITGB1', 'commot-CellChat-HSPG2-DAG1', 'commot-CellChat-PDGFA-PDGFRA', 'commot-CellChat-PDGFA-PDGFRB', 'commot-CellChat-EFNA5-EPHA4', 'commot-CellChat-EFNA5-EPHA3', 'commot-CellChat-EFNA5-EPHA7', 'commot-CellChat-MPZL1-MPZL1', 'commot-CellChat-COL1A2-SDC4', 'commot-CellChat-COL1A2-ITGA2_ITGB1', 'commot-CellChat-COL1A2-ITGA3_ITGB1', 'commot-CellChat-COL1A2-ITGA9_ITGB1', 'commot-CellChat-COL1A2-CD44', 'commot-CellChat-COL1A2-ITGAV_ITGB8', 'commot-CellChat-COL1A2-ITGA1_ITGB1', 'commot-CellChat-JAG1-NOTCH2', 'commot-CellChat-JAG1-NOTCH1', 'commot-CellChat-JAG1-NOTCH3', 'commot-CellChat-LAMB1-ITGA6_ITGB4', 'commot-CellChat-LAMB1-ITGA6_ITGB1', 'commot-CellChat-LAMB1-ITGA2_ITGB1', 'commot-CellChat-LAMB1-ITGA3_ITGB1', 'commot-CellChat-LAMB1-ITGA9_ITGB1', 'commot-CellChat-LAMB1-DAG1', 'commot-CellChat-LAMB1-CD44', 'commot-CellChat-LAMB1-ITGAV_ITGB8', 'commot-CellChat-LAMB1-ITGA1_ITGB1', 'commot-CellChat-LAMA5-ITGA6_ITGB4', 'commot-CellChat-LAMA5-ITGA6_ITGB1', 'commot-CellChat-LAMA5-ITGA2_ITGB1', 'commot-CellChat-LAMA5-ITGA3_ITGB1', 'commot-CellChat-LAMA5-ITGA9_ITGB1', 'commot-CellChat-LAMA5-DAG1', 'commot-CellChat-LAMA5-CD44', 'commot-CellChat-LAMA5-ITGAV_ITGB8', 'commot-CellChat-LAMA5-ITGA1_ITGB1', 'commot-CellChat-WNT5B-FZD6', 'commot-CellChat-GRN-SORT1', 'commot-CellChat-NRG2-ERBB2_ERBB4', 'commot-CellChat-NRG2-ERBB3', 'commot-CellChat-NRG2-ERBB4', 'commot-CellChat-NRG2-ERBB2_ERBB3', 'commot-CellChat-CSF1-CSF1R', 'commot-CellChat-F11R-JAM3', 'commot-CellChat-F11R-F11R', 'commot-CellChat-F11R-ITGAL_ITGB2', 'commot-CellChat-PECAM1-PECAM1', 'commot-CellChat-SEMA4A-NRP1_PLXNA3', 'commot-CellChat-SEMA4A-NRP1_PLXNA1', 'commot-CellChat-SEMA4A-NRP1_PLXNA2', 'commot-CellChat-SEMA4A-NRP1_PLXNA4', 'commot-CellChat-SEMA4A-PLXNB1', 'commot-CellChat-SEMA4A-PLXNB2', 'commot-CellChat-GAS6-MERTK', 'commot-CellChat-GAS6-AXL', 'commot-CellChat-PDGFC-PDGFRA', 'commot-CellChat-COL4A6-SDC4', 'commot-CellChat-COL4A6-ITGA2_ITGB1', 'commot-CellChat-COL4A6-ITGA3_ITGB1', 'commot-CellChat-COL4A6-ITGA9_ITGB1', 'commot-CellChat-COL4A6-CD44', 'commot-CellChat-COL4A6-ITGAV_ITGB8', 'commot-CellChat-COL4A6-ITGA1_ITGB1', 'commot-CellChat-LAMA4-ITGA6_ITGB4', 'commot-CellChat-LAMA4-ITGA6_ITGB1', 'commot-CellChat-LAMA4-ITGA2_ITGB1', 'commot-CellChat-LAMA4-ITGA3_ITGB1', 'commot-CellChat-LAMA4-ITGA9_ITGB1', 'commot-CellChat-LAMA4-DAG1', 'commot-CellChat-LAMA4-CD44', 'commot-CellChat-LAMA4-ITGAV_ITGB8', 'commot-CellChat-LAMA4-ITGA1_ITGB1', 'commot-CellChat-EFNB2-EPHB4', 'commot-CellChat-EFNB2-EPHA4', 'commot-CellChat-MDK-SDC2', 'commot-CellChat-MDK-ITGA4_ITGB1', 'commot-CellChat-MDK-SDC4', 'commot-CellChat-MDK-ITGA6_ITGB1', 'commot-CellChat-MDK-LRP1', 'commot-CellChat-MDK-NCL', 'commot-CellChat-COL4A3-SDC4', 'commot-CellChat-COL4A3-ITGA2_ITGB1', 'commot-CellChat-COL4A3-ITGA3_ITGB1', 'commot-CellChat-COL4A3-ITGA9_ITGB1', 'commot-CellChat-COL4A3-CD44', 'commot-CellChat-COL4A3-ITGAV_ITGB8', 'commot-CellChat-COL4A3-ITGA1_ITGB1', 'commot-CellChat-COL4A5-SDC4', 'commot-CellChat-COL4A5-ITGA2_ITGB1', 'commot-CellChat-COL4A5-ITGA3_ITGB1', 'commot-CellChat-COL4A5-ITGA9_ITGB1', 'commot-CellChat-COL4A5-CD44', 'commot-CellChat-COL4A5-ITGAV_ITGB8', 'commot-CellChat-COL4A5-ITGA1_ITGB1', 'commot-CellChat-COL6A3-SDC4', 'commot-CellChat-COL6A3-ITGA2_ITGB1', 'commot-CellChat-COL6A3-ITGA3_ITGB1', 'commot-CellChat-COL6A3-ITGA9_ITGB1', 'commot-CellChat-COL6A3-CD44', 'commot-CellChat-COL6A3-ITGAV_ITGB8', 'commot-CellChat-COL6A3-ITGA1_ITGB1', 'commot-CellChat-PTN-SDC2', 'commot-CellChat-PTN-SDC4', 'commot-CellChat-PTN-SDC3', 'commot-CellChat-PTN-NCL', 'commot-CellChat-CADM1-CADM1', 'commot-CellChat-NRG1-ERBB2_ERBB4', 'commot-CellChat-NRG1-ERBB3', 'commot-CellChat-NRG1-ERBB4', 'commot-CellChat-NRG1-ERBB2_ERBB3', 'commot-CellChat-TNC-SDC4', 'commot-CellChat-TNC-ITGA9_ITGB1', 'commot-CellChat-THBS3-CD47', 'commot-CellChat-THBS3-SDC4', 'commot-CellChat-THBS3-ITGA3_ITGB1', 'commot-CellChat-TGFB2-ACVR1_TGFBR1_TGFBR2', 'commot-CellChat-TGFB2-TGFBR1_TGFBR2', 'commot-CellChat-TGFB2-ACVR1B_TGFBR2', 'commot-CellChat-SEMA3C-NRP1_PLXNA3', 'commot-CellChat-SEMA3C-NRP1_PLXNA1', 'commot-CellChat-SEMA3C-NRP1_PLXNA2', 'commot-CellChat-SEMA3C-NRP1_PLXNA4', 'commot-CellChat-SEMA3C-PLXND1', 'commot-CellChat-TNXB-SDC4', 'commot-CellChat-TNXB-ITGA9_ITGB1', 'commot-CellChat-MIF-CD74_CXCR4', 'commot-CellChat-MIF-CD74_CD44', 'commot-CellChat-IGF1-ITGA6_ITGB4', 'commot-CellChat-IGF1-IGF1R', 'commot-CellChat-COL1A1-SDC4', 'commot-CellChat-COL1A1-ITGA2_ITGB1', 'commot-CellChat-COL1A1-ITGA3_ITGB1', 'commot-CellChat-COL1A1-ITGA9_ITGB1', 'commot-CellChat-COL1A1-CD44', 'commot-CellChat-COL1A1-ITGAV_ITGB8', 'commot-CellChat-COL1A1-ITGA1_ITGB1', 'commot-CellChat-COL4A1-SDC4', 'commot-CellChat-COL4A1-ITGA2_ITGB1', 'commot-CellChat-COL4A1-ITGA3_ITGB1', 'commot-CellChat-COL4A1-ITGA9_ITGB1', 'commot-CellChat-COL4A1-CD44', 'commot-CellChat-COL4A1-ITGAV_ITGB8', 'commot-CellChat-COL4A1-ITGA1_ITGB1', 'commot-CellChat-APP-CD74', 'commot-CellChat-CD96-NECTIN1', 'commot-CellChat-CD96-PVR', 'commot-CellChat-PTPRM-PTPRM', 'commot-CellChat-SEMA5A-PLXNA3', 'commot-CellChat-SEMA5A-PLXNA1', 'commot-CellChat-LGALS9-PTPRC', 'commot-CellChat-LGALS9-CD44', 'commot-CellChat-NEGR1-NEGR1', 'commot-CellChat-AGRN', 'commot-CellChat-APP', 'commot-CellChat-CADM', 'commot-CellChat-CD45', 'commot-CellChat-CD46', 'commot-CellChat-CD96', 'commot-CellChat-CD99', 'commot-CellChat-CDH', 'commot-CellChat-CDH1', 'commot-CellChat-COLLAGEN', 'commot-CellChat-COMPLEMENT', 'commot-CellChat-CSF', 'commot-CellChat-EPHA', 'commot-CellChat-EPHB', 'commot-CellChat-FN1', 'commot-CellChat-GALECTIN', 'commot-CellChat-GAS', 'commot-CellChat-GRN', 'commot-CellChat-HSPG', 'commot-CellChat-IGF', 'commot-CellChat-JAM', 'commot-CellChat-LAMININ', 'commot-CellChat-MIF', 'commot-CellChat-MK', 'commot-CellChat-MPZ', 'commot-CellChat-NECTIN', 'commot-CellChat-NEGR', 'commot-CellChat-NOTCH', 'commot-CellChat-NRG', 'commot-CellChat-NRXN', 'commot-CellChat-PDGF', 'commot-CellChat-PECAM1', 'commot-CellChat-PTN', 'commot-CellChat-PTPRM', 'commot-CellChat-SEMA3', 'commot-CellChat-SEMA4', 'commot-CellChat-SEMA5', 'commot-CellChat-SEMA6', 'commot-CellChat-TENASCIN', 'commot-CellChat-TGFb', 'commot-CellChat-THBS', 'commot-CellChat-VISFATIN', 'commot-CellChat-ncWNT', 'commot-CellChat-total-total'
os.environ["PATH"] += os.pathsep + '/PROJ2/FLOAT/jinwen/apps/miniconda3/envs/COMMOT/bin'
#print("total_network")
ct.pl.plot_cluster_communication_network(a_dis500, uns_names=['commot_cluster'+'-'+cluster_name+'-'+lr_database+'-total-total'],
nx_node_pos=None, nx_bg_pos=False, p_value_cutoff = 5e-2, filename='total-total_cluster.png', nx_node_cmap='Light24')
display(Image.open("./total-total_cluster.png"))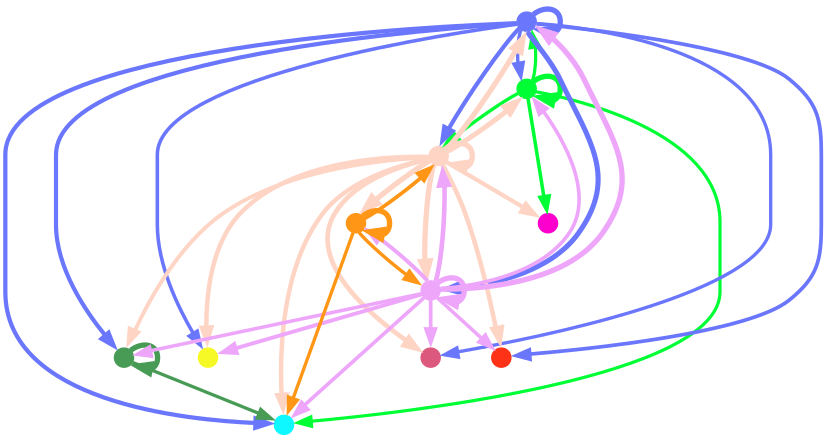
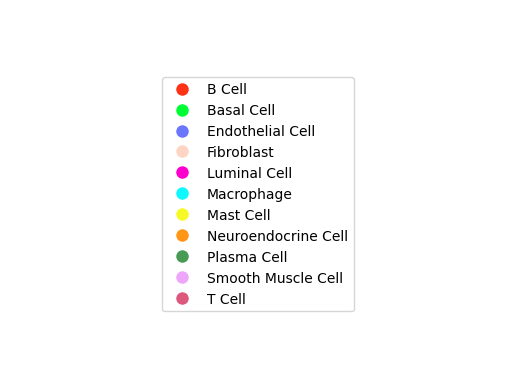
Cell communication network plot shows cell type interactions. Each color represents a cell type; lines pointing from a cell type to itself or others represent interactions.Result Files
sender_receiver_signature.png/pdf Receptor-Ligand Signal Co-expression Plot
*_signature_stream.png/pdf Cell Type Interaction Signal Flow Plot
*_cluster.png/pdf Cell Type Interaction Network Plot
*_chord_plot.png/pdf Cell Type Interaction Chord Plot Result Directory: Directory includes all images and tables involved in this analysis
Literature Case Analysis
- Literature 1: Literature "Single-cell and spatial transcriptome analyses reveal tertiary lymphoid structures linked to tumour progression and immunotherapy response in nasopharyngeal carcinoma" previously discovered CXCL13+ CAFs B cells through cell communication
References
[1] Cang Z, Zhao Y, Almet A A et al. Screening cell–cell communication in spatial transcriptomics via collective optimal transport[J]. Nat Methods, 2023, 20: 218–228.
[2] Takano Y, Suzuki J, Nomura K et al. Spatially resolved gene expression profiling of tumor microenvironment reveals key steps of lung adenocarcinoma development[J]. Nat Commun, 2024, 15: 10637.
