COMMOT 空间细胞通讯:基于最优传输算法的信号推断
背景介绍
(1)首先,使用非概率质量分布来控制运输计划的边缘,以保持物种之间的可比性;
(2)第二,对 CCC 实施空间距离约束,以避免连接空间上相距较远的部分;
(3)最后,将多物种分布(配体)传输到多物种分布(受体)以解释多物种相互作用。
COMMOT 的计算结果
%%capture
import os
import gc
import ot
import pickle
import anndata
import scanpy as sc
import pandas as pd
import numpy as np
from scipy import sparse
from scipy.stats import spearmanr, pearsonr
from scipy.spatial import distance_matrix
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.colors import Normalize
import commot as ct
import stlearn as st
from PIL import Image
import plotly
import sys
import logging
import re
import warningsTo enable them in other operations, rebuild TensorFlow with the appropriate compiler flags.
2025-09-02 09:50:38.107938: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libcudart.so.11.0'; dlerror: libcudart.so.11.0: cannot open shared object file: No such file or directory
2025-09-02 09:50:38.107975: I tensorflow/compiler/xla/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
2025-09-02 09:51:03.232851: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libnvinfer.so.7'; dlerror: libnvinfer.so.7: cannot open shared object file: No such file or directory
2025-09-02 09:51:03.234823: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libnvinfer_plugin.so.7'; dlerror: libnvinfer_plugin.so.7: cannot open shared object file: No such file or directory
2025-09-02 09:51:03.234839: W tensorflow/compiler/tf2tensorrt/utils/py_utils.cc:38] TF-TRT Warning: Cannot dlopen some TensorRT libraries. If you would like to use Nvidia GPU with TensorRT, please make sure the missing libraries mentioned above are installed properly.
COMMOT 细胞通讯模块输入参数介绍
- SAMple_name:样本名
- meta_path:注释的细胞类型文件路径,文件中需要以 barcode 为行名,列名为“celltype”(细胞类型)为列
- species:物种,与下面参数 lr_database_bool 有关,若 lr_database_bool 为 True,则该处只能填"human"或者"mouse";若为 False,则不需要填写
- lr_database:填写 COMMOT 自带的数据库名称,可选"CellChat"或者"CellPhoneDB_v4.0";若参数 provide_lr_database 是 True,则设置名称
- signaling_type:若 lr_database 填 CellChat,可选'Secreted Signaling','Cell-Cell Contact','ECM-Receptor';若 lr_database 填 CellPhoneDB_v4.0,可选'Secreted Signaling','Cell-Cell Contact';若填 None,则用选择的 lr_database 所有信号。
- col_SAM:metadata 的样本列名称
- col_celltype:metadata 的细胞类型列名称
sample_name = "N1"
meta_path = "../../data/AY1748480899609/meta.tsv"
species = "human"
lr_database = "CellChat"
signaling_type = None
col_sam="Sample"
col_celltype="CellAnnotation"
def get_cmap_qualitative(cmap_name):
if cmap_name == "Plotly":
cmap = plotly.colors.qualitative.Plotly
elif cmap_name == "Alphabet":
cmap = plotly.colors.qualitative.Alphabet
elif cmap_name == "Light24":
cmap = plotly.colors.qualitative.Light24
elif cmap_name == "Dark24":
cmap = plotly.colors.qualitative.Dark24
# Safe and Vivid are strings of form "rbg(...)"
# Handle this later.
elif cmap_name == "Safe":
cmap = plotly.colors.qualitative.Safe
elif cmap_name == "Vivid":
cmap = plotly.colors.qualitative.Vivid
return cmap- 读取 SeekSpace 数据并进行标准化、降维、聚类
adata = sc.read_10x_mtx("filtered_feature_bc_matrix/")
spatial = pd.read_csv('filtered_feature_bc_matrix/cell_locations.tsv',sep="\t",index_col=0)
spatial = spatial.loc[:,("x","y")]
selected_rows = spatial.loc[spatial.index.isin(adata.obs_names)]
selected_rows.columns = ["imagecol","imagerow"]
selected_rows = selected_rows.reindex(adata.obs_names)
selected_rows = selected_rows*0.265385
a = st.create_stlearn(count=adata.to_df(),spatial=selected_rows,library_id=sample_name, scale=1)
a.raw = a
#a.layers["raw_count"] = a.X
# Preprocessing
#st.pp.filter_genes(a,min_cells=3)
st.pp.normalize_total(a)
st.pp.log1p(a)
a_dis500 = a.copy()
# Keep raw data
#a.raw = a
st.pp.scale(a)
st.em.run_pca(a,n_comps=50,random_state=0)
st.pp.neighbors(a,n_neighbors=25,use_rep='X_pca',random_state=0)
st.tl.clustering.louvain(a,random_state=0)
sc.tl.umap(a)Log transformation step is finished in adata.X
Scale step is finished in adata.X
PCA is done! Generated in adata.obsm['X_pca'], adata.uns['pca'] and adata.varm['PCs']
Created k-Nearest-Neighbor graph in adata.uns['neighbors']
Applying Louvain cluster ...n Louvain cluster is done! The labels are stored in adata.obs['louvain']
- 将注释好的细胞类型添加到 adata 对象中
cluster_name = "celltype"
celltype = pd.read_csv(meta_path,index_col=0,sep = "\t")
celltype = celltype.loc[a.obs.index]
a.obs[cluster_name] = celltype[col_celltype]
a.obs[cluster_name] = a.obs[cluster_name].astype('category')
a_dis500 = a_dis500[a.obs.index,:]- 选择需要的受配体库,根据受配体的表达进行过滤
df_ligrec = ct.pp.ligand_receptor_database(species=species, signaling_type=signaling_type, database=lr_database)
df_ligrec_filtered = ct.pp.filter_lr_database(df_ligrec, a_dis500, min_cell_pct=0.05)
df_ligrec_filtered| 0 | 1 | 2 | 3 | |
|---|---|---|---|---|
| 0 | TGFB1 | TGFBR1_TGFBR2 | TGFb | Secreted Signaling |
| 1 | TGFB2 | TGFBR1_TGFBR2 | TGFb | Secreted Signaling |
| 2 | TGFB3 | TGFBR1_TGFBR2 | TGFb | Secreted Signaling |
| 3 | TGFB1 | ACVR1B_TGFBR2 | TGFb | Secreted Signaling |
| 4 | TGFB2 | ACVR1B_TGFBR2 | TGFb | Secreted Signaling |
| ... | ... | ... | ... | ... |
| 252 | SEMA4G | PLXNB2 | SEMA4 | Cell-Cell Contact |
| 253 | SEMA5A | PLXNA1 | SEMA5 | Cell-Cell Contact |
| 254 | SEMA5A | PLXNA3 | SEMA5 | Cell-Cell Contact |
| 255 | SEMA6A | PLXNA2 | SEMA6 | Cell-Cell Contact |
| 256 | SEMA6A | PLXNA4 | SEMA6 | Cell-Cell Contact |
257 rows × 4 columns
- 针对过滤后的受配体进行细胞通讯推断,距离范围在 500um。
ct.tl.spatial_communication(a_dis500,
database_name=lr_database, df_ligrec=df_ligrec_filtered, dis_thr=500, heteromeric=True, pathway_sum=True)a_dis500.write("./adata_dis500.h5ad")受配体对发送和接收信号的数量
pts = a_dis500.obsm['spatial']
sender = a_dis500.obsm['commot-' + lr_database + '-sum-sender'][a_dis500.obsm['commot-' + lr_database + '-sum-sender'].columns[1]]
receiver = a_dis500.obsm['commot-' + lr_database + '-sum-receiver'][a_dis500.obsm['commot-' + lr_database + '-sum-receiver'].columns[1]]
fig, ax = plt.subplots(1,2, figsize=(20,8))
ax[0].scatter(pts[:,0], pts[:,1], c=sender, s=5, cmap='Blues')
ax[0].set_title(a_dis500.obsm['commot-' + lr_database + '-sum-sender'].columns[1]+'_Sender')
norm_sender = Normalize(vmin=min(sender), vmax=max(sender))
plt.colorbar(cm.ScalarMappable(norm=norm_sender, cmap='Blues'), ax=ax[0])
ax[0].set_aspect("equal")
ax[1].scatter(pts[:,0], pts[:,1], c=receiver, s=5, cmap='Reds')
ax[1].set_title(a_dis500.obsm['commot-' + lr_database + '-sum-receiver'].columns[1]+'_Receiver')
norm_receiver = Normalize(vmin=min(receiver), vmax=max(receiver))
plt.colorbar(cm.ScalarMappable(norm=norm_receiver, cmap='Reds'), ax=ax[1])
ax[1].set_aspect("equal")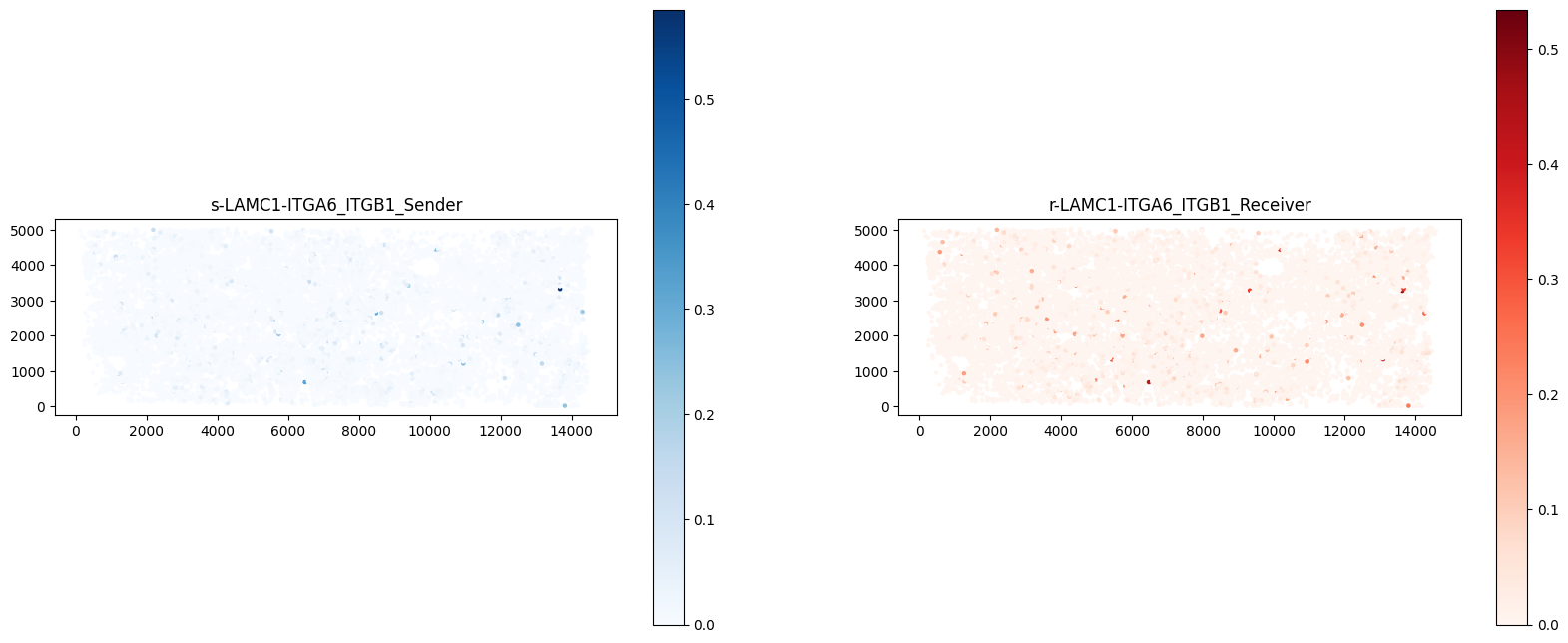
受配体共表达情况。左图代表着配体的表达情况;右图代表着受体的表达情况;颜色越深表达越高。若配受体表达在同一区域,从而推断出表达该受配体细胞之间会发生互作。(这里以一种受配体对展示)#ct.tl.communication_direction(a_dis500, database_name=lr_database, pathway_name='PSAP', k=5)
#ct.pl.plot_cell_communication(a_dis500, database_name=lr_database, pathway_name='PSAP', plot_method='grid', background_legend=True,
# scale=0.00003, ndsize=8, grid_density=0.4, summary='sender', background='image', clustering=cluster_name, cmap='Alphabet',
# normalize_v = True, normalize_v_quantile=0.995)细胞通讯信号流可视化
COMMOT 认为细胞通讯是某个细胞会与邻近细胞互作,邻近细胞与下一个邻近细胞互作,依次进行就会形成“信号流”,图中展示了通路的信号流方向。
#print("total_signature_stream")
ct.tl.communication_direction(a_dis500, database_name=lr_database, k=5)
ct.pl.plot_cell_communication(a_dis500, database_name=lr_database, lr_pair=('total','total'), plot_method='stream', background_legend=True,
scale=0.00003, ndsize=8, grid_density=0.4, summary='sender', background='summary', clustering=cluster_name, cmap='Reds',
normalize_v = True, normalize_v_quantile=0.995)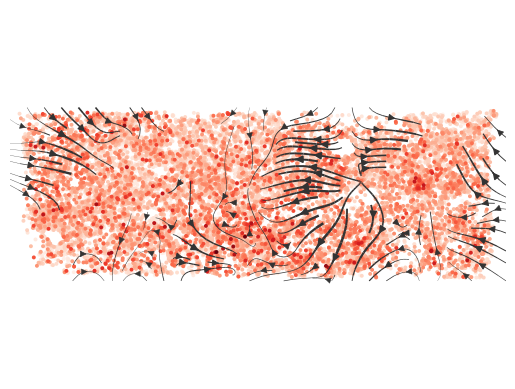
每个细胞之间通讯的方向性。箭头代表着通讯的方向,底色代表着细胞发送信号的强度。- 计算不同细胞类型之间通讯交流强度
a_dis500.obs[cluster_name] = a.obs[cluster_name]
#计算每个受配体对的结果
for i,j in enumerate(df_ligrec_filtered.values):
ct.tl.cluster_communication(a_dis500, database_name=lr_database, clustering=cluster_name,n_permutations=100,lr_pair=(j[0],j[1]))
ct.tl.cluster_communication(a_dis500, database_name=lr_database, clustering=cluster_name,n_permutations=100)细胞通讯网络
a_dis500obs: 'imagecol', 'imagerow', 'celltype'
uns: 'spatial', 'log1p', 'commot-CellChat-info', 'commot_cluster-celltype-CellChat-TGFB1-TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB2-TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB3-TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB1-ACVR1B_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB2-ACVR1B_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB3-ACVR1B_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB1-ACVR1_TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB2-ACVR1_TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-TGFB3-ACVR1_TGFBR1_TGFBR2', 'commot_cluster-celltype-CellChat-WNT5B-FZD6', 'commot_cluster-celltype-CellChat-NRG1-ERBB3', 'commot_cluster-celltype-CellChat-NRG1-ERBB2_ERBB3', 'commot_cluster-celltype-CellChat-NRG1-ERBB4', 'commot_cluster-celltype-CellChat-NRG1-ERBB2_ERBB4', 'commot_cluster-celltype-CellChat-NRG2-ERBB3', 'commot_cluster-celltype-CellChat-NRG2-ERBB2_ERBB3', 'commot_cluster-celltype-CellChat-NRG2-ERBB4', 'commot_cluster-celltype-CellChat-NRG2-ERBB2_ERBB4', 'commot_cluster-celltype-CellChat-PDGFA-PDGFRA', 'commot_cluster-celltype-CellChat-PDGFA-PDGFRB', 'commot_cluster-celltype-CellChat-PDGFC-PDGFRA', 'commot_cluster-celltype-CellChat-IGF1-IGF1R', 'commot_cluster-celltype-CellChat-IGF1-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-MIF-CD74_CXCR4', 'commot_cluster-celltype-CellChat-MIF-CD74_CD44', 'commot_cluster-celltype-CellChat-CSF1-CSF1R', 'commot_cluster-celltype-CellChat-NAMPT-INSR', 'commot_cluster-celltype-CellChat-NAMPT-ITGA5_ITGB1', 'commot_cluster-celltype-CellChat-MDK-SDC2', 'commot_cluster-celltype-CellChat-MDK-SDC4', 'commot_cluster-celltype-CellChat-MDK-ITGA4_ITGB1', 'commot_cluster-celltype-CellChat-MDK-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-MDK-LRP1', 'commot_cluster-celltype-CellChat-MDK-NCL', 'commot_cluster-celltype-CellChat-PTN-SDC2', 'commot_cluster-celltype-CellChat-PTN-SDC3', 'commot_cluster-celltype-CellChat-PTN-SDC4', 'commot_cluster-celltype-CellChat-PTN-NCL', 'commot_cluster-celltype-CellChat-C3-ITGAX_ITGB2', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA1', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA2', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA3', 'commot_cluster-celltype-CellChat-SEMA3C-NRP1_PLXNA4', 'commot_cluster-celltype-CellChat-SEMA3C-PLXND1', 'commot_cluster-celltype-CellChat-GAS6-AXL', 'commot_cluster-celltype-CellChat-GAS6-MERTK', 'commot_cluster-celltype-CellChat-GRN-SORT1', 'commot_cluster-celltype-CellChat-LGALS9-PTPRC', 'commot_cluster-celltype-CellChat-LGALS9-CD44', 'commot_cluster-celltype-CellChat-COL1A1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-COL1A1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGA4_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGA5_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA1_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-THBS1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-THBS3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL1A1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA3_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA6_ITGB1', 'commot_cluster-celltype-CellChat-TNC-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-TNXB-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL1A1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL1A2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A3-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A4-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A5-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL4A6-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL6A1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL6A2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-COL6A3-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA3-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA4-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMA5-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMB1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMB2-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-LAMC1-ITGA9_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGAV_ITGB1', 'commot_cluster-celltype-CellChat-FN1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA3-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA4-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA5-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMB1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMB2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMC1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL1A1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL1A2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A3-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A4-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A5-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL4A6-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL6A1-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL6A2-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-COL6A3-ITGAV_ITGB8', 'commot_cluster-celltype-CellChat-LAMA2-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMA3-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMA4-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMA5-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMB1-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMB2-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-LAMC1-ITGA6_ITGB4', 'commot_cluster-celltype-CellChat-FN1-CD44', 'commot_cluster-celltype-CellChat-COL1A1-CD44', 'commot_cluster-celltype-CellChat-COL1A2-CD44', 'commot_cluster-celltype-CellChat-COL4A1-CD44', 'commot_cluster-celltype-CellChat-COL4A2-CD44', 'commot_cluster-celltype-CellChat-COL4A3-CD44', 'commot_cluster-celltype-CellChat-COL4A4-CD44', 'commot_cluster-celltype-CellChat-COL4A5-CD44', 'commot_cluster-celltype-CellChat-COL4A6-CD44', 'commot_cluster-celltype-CellChat-COL6A1-CD44', 'commot_cluster-celltype-CellChat-COL6A2-CD44', 'commot_cluster-celltype-CellChat-COL6A3-CD44', 'commot_cluster-celltype-CellChat-LAMA2-CD44', 'commot_cluster-celltype-CellChat-LAMA3-CD44', 'commot_cluster-celltype-CellChat-LAMA4-CD44', 'commot_cluster-celltype-CellChat-LAMA5-CD44', 'commot_cluster-celltype-CellChat-LAMB1-CD44', 'commot_cluster-celltype-CellChat-LAMB2-CD44', 'commot_cluster-celltype-CellChat-LAMC1-CD44', 'commot_cluster-celltype-CellChat-COL1A1-SDC4', 'commot_cluster-celltype-CellChat-COL1A2-SDC4', 'commot_cluster-celltype-CellChat-COL4A1-SDC4', 'commot_cluster-celltype-CellChat-COL4A2-SDC4', 'commot_cluster-celltype-CellChat-COL4A3-SDC4', 'commot_cluster-celltype-CellChat-COL4A4-SDC4', 'commot_cluster-celltype-CellChat-COL4A5-SDC4', 'commot_cluster-celltype-CellChat-COL4A6-SDC4', 'commot_cluster-celltype-CellChat-COL6A1-SDC4', 'commot_cluster-celltype-CellChat-COL6A2-SDC4', 'commot_cluster-celltype-CellChat-COL6A3-SDC4', 'commot_cluster-celltype-CellChat-FN1-SDC4', 'commot_cluster-celltype-CellChat-TNC-SDC4', 'commot_cluster-celltype-CellChat-TNXB-SDC4', 'commot_cluster-celltype-CellChat-THBS1-SDC4', 'commot_cluster-celltype-CellChat-THBS3-SDC4', 'commot_cluster-celltype-CellChat-THBS1-CD47', 'commot_cluster-celltype-CellChat-THBS3-CD47', 'commot_cluster-celltype-CellChat-AGRN-DAG1', 'commot_cluster-celltype-CellChat-HSPG2-DAG1', 'commot_cluster-celltype-CellChat-LAMA2-DAG1', 'commot_cluster-celltype-CellChat-LAMA3-DAG1', 'commot_cluster-celltype-CellChat-LAMA4-DAG1', 'commot_cluster-celltype-CellChat-LAMA5-DAG1', 'commot_cluster-celltype-CellChat-LAMB1-DAG1', 'commot_cluster-celltype-CellChat-LAMB2-DAG1', 'commot_cluster-celltype-CellChat-LAMC1-DAG1', 'commot_cluster-celltype-CellChat-APP-CD74', 'commot_cluster-celltype-CellChat-CADM1-CADM1', 'commot_cluster-celltype-CellChat-PTPRC-MRC1', 'commot_cluster-celltype-CellChat-CD46-JAG1', 'commot_cluster-celltype-CellChat-CD96-NECTIN1', 'commot_cluster-celltype-CellChat-CD96-PVR', 'commot_cluster-celltype-CellChat-CD99-CD99', 'commot_cluster-celltype-CellChat-CD99-CD99L2', 'commot_cluster-celltype-CellChat-CDH1-CDH1', 'commot_cluster-celltype-CellChat-CDH1-ITGA2_ITGB1', 'commot_cluster-celltype-CellChat-EFNA1-EPHA3', 'commot_cluster-celltype-CellChat-EFNA1-EPHA4', 'commot_cluster-celltype-CellChat-EFNA1-EPHA7', 'commot_cluster-celltype-CellChat-EFNA5-EPHA3', 'commot_cluster-celltype-CellChat-EFNA5-EPHA4', 'commot_cluster-celltype-CellChat-EFNA5-EPHA7', 'commot_cluster-celltype-CellChat-EFNB2-EPHA4', 'commot_cluster-celltype-CellChat-EFNB2-EPHB4', 'commot_cluster-celltype-CellChat-F11R-ITGAL_ITGB2', 'commot_cluster-celltype-CellChat-F11R-F11R', 'commot_cluster-celltype-CellChat-F11R-JAM3', 'commot_cluster-celltype-CellChat-JAM3-F11R', 'commot_cluster-celltype-CellChat-JAM3-JAM3', 'commot_cluster-celltype-CellChat-MPZL1-MPZL1', 'commot_cluster-celltype-CellChat-NECTIN1-CD96', 'commot_cluster-celltype-CellChat-NEGR1-NEGR1', 'commot_cluster-celltype-CellChat-DLL1-NOTCH1', 'commot_cluster-celltype-CellChat-DLL1-NOTCH3', 'commot_cluster-celltype-CellChat-JAG1-NOTCH1', 'commot_cluster-celltype-CellChat-JAG1-NOTCH2', 'commot_cluster-celltype-CellChat-JAG1-NOTCH3', 'commot_cluster-celltype-CellChat-DLL1-NOTCH2', 'commot_cluster-celltype-CellChat-NRXN3-NLGN1', 'commot_cluster-celltype-CellChat-NRXN3-NLGN2', 'commot_cluster-celltype-CellChat-PECAM1-PECAM1', 'commot_cluster-celltype-CellChat-PTPRM-PTPRM', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA1', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA2', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA3', 'commot_cluster-celltype-CellChat-SEMA4A-NRP1_PLXNA4', 'commot_cluster-celltype-CellChat-SEMA4A-PLXNB1', 'commot_cluster-celltype-CellChat-SEMA4A-PLXNB2', 'commot_cluster-celltype-CellChat-SEMA4D-PLXNB1', 'commot_cluster-celltype-CellChat-SEMA4D-PLXNB2', 'commot_cluster-celltype-CellChat-SEMA4G-PLXNB2', 'commot_cluster-celltype-CellChat-SEMA5A-PLXNA1', 'commot_cluster-celltype-CellChat-SEMA5A-PLXNA3', 'commot_cluster-celltype-CellChat-SEMA6A-PLXNA2', 'commot_cluster-celltype-CellChat-SEMA6A-PLXNA4', 'commot_cluster-celltype-CellChat-total-total'
obsm: 'spatial', 'commot-CellChat-sum-sender', 'commot-CellChat-sum-receiver', 'commot_sender_vf-CellChat-total-total', 'commot_receiver_vf-CellChat-total-total'
obsp: 'commot-CellChat-LAMC1-ITGA6_ITGB4', 'commot-CellChat-LAMC1-ITGA6_ITGB1', 'commot-CellChat-LAMC1-ITGA2_ITGB1', 'commot-CellChat-LAMC1-ITGA3_ITGB1', 'commot-CellChat-LAMC1-ITGA9_ITGB1', 'commot-CellChat-LAMC1-DAG1', 'commot-CellChat-LAMC1-CD44', 'commot-CellChat-LAMC1-ITGAV_ITGB8', 'commot-CellChat-LAMC1-ITGA1_ITGB1', 'commot-CellChat-LAMB2-ITGA6_ITGB4', 'commot-CellChat-LAMB2-ITGA6_ITGB1', 'commot-CellChat-LAMB2-ITGA2_ITGB1', 'commot-CellChat-LAMB2-ITGA3_ITGB1', 'commot-CellChat-LAMB2-ITGA9_ITGB1', 'commot-CellChat-LAMB2-DAG1', 'commot-CellChat-LAMB2-CD44', 'commot-CellChat-LAMB2-ITGAV_ITGB8', 'commot-CellChat-LAMB2-ITGA1_ITGB1', 'commot-CellChat-NAMPT-INSR', 'commot-CellChat-NAMPT-ITGA5_ITGB1', 'commot-CellChat-COL6A1-SDC4', 'commot-CellChat-COL6A1-ITGA2_ITGB1', 'commot-CellChat-COL6A1-ITGA3_ITGB1', 'commot-CellChat-COL6A1-ITGA9_ITGB1', 'commot-CellChat-COL6A1-CD44', 'commot-CellChat-COL6A1-ITGAV_ITGB8', 'commot-CellChat-COL6A1-ITGA1_ITGB1', 'commot-CellChat-THBS1-CD47', 'commot-CellChat-THBS1-SDC4', 'commot-CellChat-THBS1-ITGA3_ITGB1', 'commot-CellChat-JAM3-JAM3', 'commot-CellChat-JAM3-F11R', 'commot-CellChat-SEMA4G-PLXNB2', 'commot-CellChat-NRXN3-NLGN2', 'commot-CellChat-NRXN3-NLGN1', 'commot-CellChat-COL4A4-SDC4', 'commot-CellChat-COL4A4-ITGA2_ITGB1', 'commot-CellChat-COL4A4-ITGA3_ITGB1', 'commot-CellChat-COL4A4-ITGA9_ITGB1', 'commot-CellChat-COL4A4-CD44', 'commot-CellChat-COL4A4-ITGAV_ITGB8', 'commot-CellChat-COL4A4-ITGA1_ITGB1', 'commot-CellChat-PTPRC-MRC1', 'commot-CellChat-CD99-CD99L2', 'commot-CellChat-CD99-CD99', 'commot-CellChat-TGFB1-ACVR1_TGFBR1_TGFBR2', 'commot-CellChat-TGFB1-TGFBR1_TGFBR2', 'commot-CellChat-TGFB1-ACVR1B_TGFBR2', 'commot-CellChat-COL4A2-SDC4', 'commot-CellChat-COL4A2-ITGA2_ITGB1', 'commot-CellChat-COL4A2-ITGA3_ITGB1', 'commot-CellChat-COL4A2-ITGA9_ITGB1', 'commot-CellChat-COL4A2-CD44', 'commot-CellChat-COL4A2-ITGAV_ITGB8', 'commot-CellChat-COL4A2-ITGA1_ITGB1', 'commot-CellChat-TGFB3-ACVR1_TGFBR1_TGFBR2', 'commot-CellChat-TGFB3-TGFBR1_TGFBR2', 'commot-CellChat-TGFB3-ACVR1B_TGFBR2', 'commot-CellChat-LAMA3-ITGA6_ITGB4', 'commot-CellChat-LAMA3-ITGA6_ITGB1', 'commot-CellChat-LAMA3-ITGA2_ITGB1', 'commot-CellChat-LAMA3-ITGA3_ITGB1', 'commot-CellChat-LAMA3-ITGA9_ITGB1', 'commot-CellChat-LAMA3-DAG1', 'commot-CellChat-LAMA3-CD44', 'commot-CellChat-LAMA3-ITGAV_ITGB8', 'commot-CellChat-LAMA3-ITGA1_ITGB1', 'commot-CellChat-CDH1-ITGA2_ITGB1', 'commot-CellChat-CDH1-CDH1', 'commot-CellChat-LAMA2-ITGA6_ITGB4', 'commot-CellChat-LAMA2-ITGA6_ITGB1', 'commot-CellChat-LAMA2-ITGA2_ITGB1', 'commot-CellChat-LAMA2-ITGA3_ITGB1', 'commot-CellChat-LAMA2-ITGA9_ITGB1', 'commot-CellChat-LAMA2-DAG1', 'commot-CellChat-LAMA2-CD44', 'commot-CellChat-LAMA2-ITGAV_ITGB8', 'commot-CellChat-LAMA2-ITGA1_ITGB1', 'commot-CellChat-EFNA1-EPHA4', 'commot-CellChat-EFNA1-EPHA3', 'commot-CellChat-EFNA1-EPHA7', 'commot-CellChat-CD46-JAG1', 'commot-CellChat-AGRN-DAG1', 'commot-CellChat-C3-ITGAX_ITGB2', 'commot-CellChat-NECTIN1-CD96', 'commot-CellChat-DLL1-NOTCH2', 'commot-CellChat-DLL1-NOTCH1', 'commot-CellChat-DLL1-NOTCH3', 'commot-CellChat-SEMA4D-PLXNB1', 'commot-CellChat-SEMA4D-PLXNB2', 'commot-CellChat-SEMA6A-PLXNA2', 'commot-CellChat-SEMA6A-PLXNA4', 'commot-CellChat-COL6A2-SDC4', 'commot-CellChat-COL6A2-ITGA2_ITGB1', 'commot-CellChat-COL6A2-ITGA3_ITGB1', 'commot-CellChat-COL6A2-ITGA9_ITGB1', 'commot-CellChat-COL6A2-CD44', 'commot-CellChat-COL6A2-ITGAV_ITGB8', 'commot-CellChat-COL6A2-ITGA1_ITGB1', 'commot-CellChat-FN1-ITGA4_ITGB1', 'commot-CellChat-FN1-SDC4', 'commot-CellChat-FN1-ITGA3_ITGB1', 'commot-CellChat-FN1-CD44', 'commot-CellChat-FN1-ITGAV_ITGB8', 'commot-CellChat-FN1-ITGA5_ITGB1', 'commot-CellChat-FN1-ITGAV_ITGB1', 'commot-CellChat-HSPG2-DAG1', 'commot-CellChat-PDGFA-PDGFRA', 'commot-CellChat-PDGFA-PDGFRB', 'commot-CellChat-EFNA5-EPHA4', 'commot-CellChat-EFNA5-EPHA3', 'commot-CellChat-EFNA5-EPHA7', 'commot-CellChat-MPZL1-MPZL1', 'commot-CellChat-COL1A2-SDC4', 'commot-CellChat-COL1A2-ITGA2_ITGB1', 'commot-CellChat-COL1A2-ITGA3_ITGB1', 'commot-CellChat-COL1A2-ITGA9_ITGB1', 'commot-CellChat-COL1A2-CD44', 'commot-CellChat-COL1A2-ITGAV_ITGB8', 'commot-CellChat-COL1A2-ITGA1_ITGB1', 'commot-CellChat-JAG1-NOTCH2', 'commot-CellChat-JAG1-NOTCH1', 'commot-CellChat-JAG1-NOTCH3', 'commot-CellChat-LAMB1-ITGA6_ITGB4', 'commot-CellChat-LAMB1-ITGA6_ITGB1', 'commot-CellChat-LAMB1-ITGA2_ITGB1', 'commot-CellChat-LAMB1-ITGA3_ITGB1', 'commot-CellChat-LAMB1-ITGA9_ITGB1', 'commot-CellChat-LAMB1-DAG1', 'commot-CellChat-LAMB1-CD44', 'commot-CellChat-LAMB1-ITGAV_ITGB8', 'commot-CellChat-LAMB1-ITGA1_ITGB1', 'commot-CellChat-LAMA5-ITGA6_ITGB4', 'commot-CellChat-LAMA5-ITGA6_ITGB1', 'commot-CellChat-LAMA5-ITGA2_ITGB1', 'commot-CellChat-LAMA5-ITGA3_ITGB1', 'commot-CellChat-LAMA5-ITGA9_ITGB1', 'commot-CellChat-LAMA5-DAG1', 'commot-CellChat-LAMA5-CD44', 'commot-CellChat-LAMA5-ITGAV_ITGB8', 'commot-CellChat-LAMA5-ITGA1_ITGB1', 'commot-CellChat-WNT5B-FZD6', 'commot-CellChat-GRN-SORT1', 'commot-CellChat-NRG2-ERBB2_ERBB4', 'commot-CellChat-NRG2-ERBB3', 'commot-CellChat-NRG2-ERBB4', 'commot-CellChat-NRG2-ERBB2_ERBB3', 'commot-CellChat-CSF1-CSF1R', 'commot-CellChat-F11R-JAM3', 'commot-CellChat-F11R-F11R', 'commot-CellChat-F11R-ITGAL_ITGB2', 'commot-CellChat-PECAM1-PECAM1', 'commot-CellChat-SEMA4A-NRP1_PLXNA3', 'commot-CellChat-SEMA4A-NRP1_PLXNA1', 'commot-CellChat-SEMA4A-NRP1_PLXNA2', 'commot-CellChat-SEMA4A-NRP1_PLXNA4', 'commot-CellChat-SEMA4A-PLXNB1', 'commot-CellChat-SEMA4A-PLXNB2', 'commot-CellChat-GAS6-MERTK', 'commot-CellChat-GAS6-AXL', 'commot-CellChat-PDGFC-PDGFRA', 'commot-CellChat-COL4A6-SDC4', 'commot-CellChat-COL4A6-ITGA2_ITGB1', 'commot-CellChat-COL4A6-ITGA3_ITGB1', 'commot-CellChat-COL4A6-ITGA9_ITGB1', 'commot-CellChat-COL4A6-CD44', 'commot-CellChat-COL4A6-ITGAV_ITGB8', 'commot-CellChat-COL4A6-ITGA1_ITGB1', 'commot-CellChat-LAMA4-ITGA6_ITGB4', 'commot-CellChat-LAMA4-ITGA6_ITGB1', 'commot-CellChat-LAMA4-ITGA2_ITGB1', 'commot-CellChat-LAMA4-ITGA3_ITGB1', 'commot-CellChat-LAMA4-ITGA9_ITGB1', 'commot-CellChat-LAMA4-DAG1', 'commot-CellChat-LAMA4-CD44', 'commot-CellChat-LAMA4-ITGAV_ITGB8', 'commot-CellChat-LAMA4-ITGA1_ITGB1', 'commot-CellChat-EFNB2-EPHB4', 'commot-CellChat-EFNB2-EPHA4', 'commot-CellChat-MDK-SDC2', 'commot-CellChat-MDK-ITGA4_ITGB1', 'commot-CellChat-MDK-SDC4', 'commot-CellChat-MDK-ITGA6_ITGB1', 'commot-CellChat-MDK-LRP1', 'commot-CellChat-MDK-NCL', 'commot-CellChat-COL4A3-SDC4', 'commot-CellChat-COL4A3-ITGA2_ITGB1', 'commot-CellChat-COL4A3-ITGA3_ITGB1', 'commot-CellChat-COL4A3-ITGA9_ITGB1', 'commot-CellChat-COL4A3-CD44', 'commot-CellChat-COL4A3-ITGAV_ITGB8', 'commot-CellChat-COL4A3-ITGA1_ITGB1', 'commot-CellChat-COL4A5-SDC4', 'commot-CellChat-COL4A5-ITGA2_ITGB1', 'commot-CellChat-COL4A5-ITGA3_ITGB1', 'commot-CellChat-COL4A5-ITGA9_ITGB1', 'commot-CellChat-COL4A5-CD44', 'commot-CellChat-COL4A5-ITGAV_ITGB8', 'commot-CellChat-COL4A5-ITGA1_ITGB1', 'commot-CellChat-COL6A3-SDC4', 'commot-CellChat-COL6A3-ITGA2_ITGB1', 'commot-CellChat-COL6A3-ITGA3_ITGB1', 'commot-CellChat-COL6A3-ITGA9_ITGB1', 'commot-CellChat-COL6A3-CD44', 'commot-CellChat-COL6A3-ITGAV_ITGB8', 'commot-CellChat-COL6A3-ITGA1_ITGB1', 'commot-CellChat-PTN-SDC2', 'commot-CellChat-PTN-SDC4', 'commot-CellChat-PTN-SDC3', 'commot-CellChat-PTN-NCL', 'commot-CellChat-CADM1-CADM1', 'commot-CellChat-NRG1-ERBB2_ERBB4', 'commot-CellChat-NRG1-ERBB3', 'commot-CellChat-NRG1-ERBB4', 'commot-CellChat-NRG1-ERBB2_ERBB3', 'commot-CellChat-TNC-SDC4', 'commot-CellChat-TNC-ITGA9_ITGB1', 'commot-CellChat-THBS3-CD47', 'commot-CellChat-THBS3-SDC4', 'commot-CellChat-THBS3-ITGA3_ITGB1', 'commot-CellChat-TGFB2-ACVR1_TGFBR1_TGFBR2', 'commot-CellChat-TGFB2-TGFBR1_TGFBR2', 'commot-CellChat-TGFB2-ACVR1B_TGFBR2', 'commot-CellChat-SEMA3C-NRP1_PLXNA3', 'commot-CellChat-SEMA3C-NRP1_PLXNA1', 'commot-CellChat-SEMA3C-NRP1_PLXNA2', 'commot-CellChat-SEMA3C-NRP1_PLXNA4', 'commot-CellChat-SEMA3C-PLXND1', 'commot-CellChat-TNXB-SDC4', 'commot-CellChat-TNXB-ITGA9_ITGB1', 'commot-CellChat-MIF-CD74_CXCR4', 'commot-CellChat-MIF-CD74_CD44', 'commot-CellChat-IGF1-ITGA6_ITGB4', 'commot-CellChat-IGF1-IGF1R', 'commot-CellChat-COL1A1-SDC4', 'commot-CellChat-COL1A1-ITGA2_ITGB1', 'commot-CellChat-COL1A1-ITGA3_ITGB1', 'commot-CellChat-COL1A1-ITGA9_ITGB1', 'commot-CellChat-COL1A1-CD44', 'commot-CellChat-COL1A1-ITGAV_ITGB8', 'commot-CellChat-COL1A1-ITGA1_ITGB1', 'commot-CellChat-COL4A1-SDC4', 'commot-CellChat-COL4A1-ITGA2_ITGB1', 'commot-CellChat-COL4A1-ITGA3_ITGB1', 'commot-CellChat-COL4A1-ITGA9_ITGB1', 'commot-CellChat-COL4A1-CD44', 'commot-CellChat-COL4A1-ITGAV_ITGB8', 'commot-CellChat-COL4A1-ITGA1_ITGB1', 'commot-CellChat-APP-CD74', 'commot-CellChat-CD96-NECTIN1', 'commot-CellChat-CD96-PVR', 'commot-CellChat-PTPRM-PTPRM', 'commot-CellChat-SEMA5A-PLXNA3', 'commot-CellChat-SEMA5A-PLXNA1', 'commot-CellChat-LGALS9-PTPRC', 'commot-CellChat-LGALS9-CD44', 'commot-CellChat-NEGR1-NEGR1', 'commot-CellChat-AGRN', 'commot-CellChat-APP', 'commot-CellChat-CADM', 'commot-CellChat-CD45', 'commot-CellChat-CD46', 'commot-CellChat-CD96', 'commot-CellChat-CD99', 'commot-CellChat-CDH', 'commot-CellChat-CDH1', 'commot-CellChat-COLLAGEN', 'commot-CellChat-COMPLEMENT', 'commot-CellChat-CSF', 'commot-CellChat-EPHA', 'commot-CellChat-EPHB', 'commot-CellChat-FN1', 'commot-CellChat-GALECTIN', 'commot-CellChat-GAS', 'commot-CellChat-GRN', 'commot-CellChat-HSPG', 'commot-CellChat-IGF', 'commot-CellChat-JAM', 'commot-CellChat-LAMININ', 'commot-CellChat-MIF', 'commot-CellChat-MK', 'commot-CellChat-MPZ', 'commot-CellChat-NECTIN', 'commot-CellChat-NEGR', 'commot-CellChat-NOTCH', 'commot-CellChat-NRG', 'commot-CellChat-NRXN', 'commot-CellChat-PDGF', 'commot-CellChat-PECAM1', 'commot-CellChat-PTN', 'commot-CellChat-PTPRM', 'commot-CellChat-SEMA3', 'commot-CellChat-SEMA4', 'commot-CellChat-SEMA5', 'commot-CellChat-SEMA6', 'commot-CellChat-TENASCIN', 'commot-CellChat-TGFb', 'commot-CellChat-THBS', 'commot-CellChat-VISFATIN', 'commot-CellChat-ncWNT', 'commot-CellChat-total-total'
os.environ["PATH"] += os.pathsep + '/PROJ2/FLOAT/jinwen/apps/miniconda3/envs/COMMOT/bin'
#print("total_network")
ct.pl.plot_cluster_communication_network(a_dis500, uns_names=['commot_cluster'+'-'+cluster_name+'-'+lr_database+'-total-total'],
nx_node_pos=None, nx_bg_pos=False, p_value_cutoff = 5e-2, filename='total-total_cluster.png', nx_node_cmap='Light24')
display(Image.open("./total-total_cluster.png"))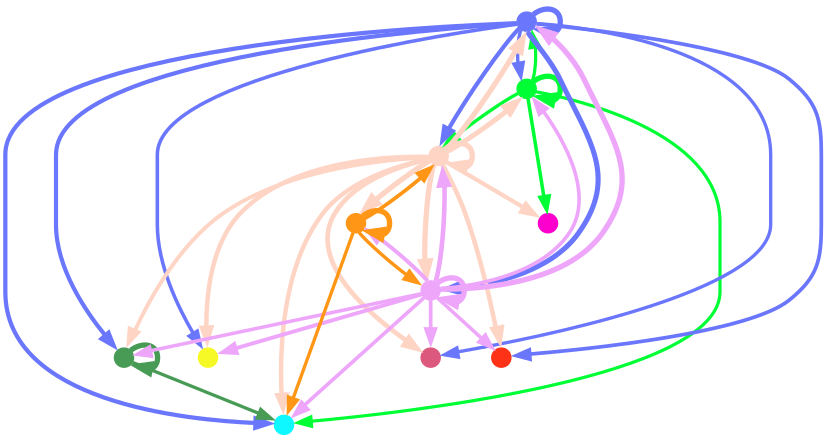
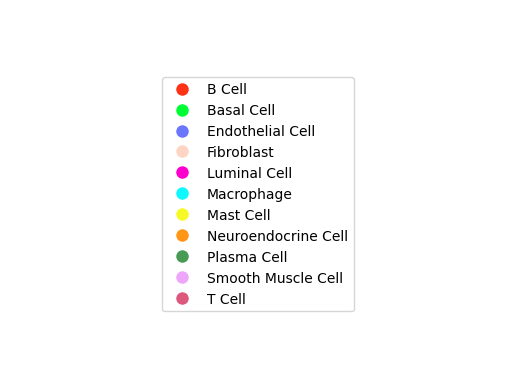
细胞通讯网络图中展示细胞类型互作情况。每一种颜色代表着一种细胞类型;线条从一个细胞类型指向自己或者其他的细胞类型,代表着互作的情况。结果文件
sender_receiver_signature.png/pdf 受配体信号共表达图
*_signature_stream.png/pdf 细胞类型互作信号流图
*_cluster.png/pdf 细胞类型互作网络图
*_chord_plot.png/pdf 细胞类型互作和弦图 结果目录:目录下包括此项分析所涉及的所有图片以及表格等结果文件
文献案例解析
- 文献一: 文献《Single-cell and spatial transcriptome analyses reveal tertiary lymphoid structures linked to tumour progression and immunotherapy response in nasopharyngeal carcinoma》前文通过细胞通讯发现 CXCL13+ CAFs B cells
参考资料
[1] Cang Z, Zhao Y, Almet A A et al. Screening cell–cell communication in spatial transcriptomics via collective optimal transport[J]. Nat Methods, 2023, 20: 218–228.
[2] Takano Y, Suzuki J, Nomura K et al. Spatially resolved gene expression profiling of tumor microenvironment reveals key steps of lung adenocarcinoma development[J]. Nat Commun, 2024, 15: 10637.
