单细胞空间微环境解析:细胞 Niche 分析与聚类教程
时长: 6 分钟
字数: 1.1k 字
更新: 2026-02-28
阅读: 0 次
加载分析包
python
import numpy as np
import pandas as pd
import scanpy as sc
import seaborn as sns
from sklearn.neighbors import NearestNeighbors
from sklearn.cluster import MiniBatchKMeans
import matplotlib.pyplot as plt
from matplotlib.pyplot import rc_context
import stlearn as stoutput
/PROJ2/FLOAT/jinwen/apps/miniconda3/envs/COMMOT/lib/python3.10/site-packages/stlearn/tools/microenv/cci/het.py:192: NumbaDeprecationWarning: The keyword argument 'nopython=False' was supplied. From Numba 0.59.0 the default is being changed to True and use of 'nopython=False' will raise a warning as the argument will have no effect. See https://numba.readthedocs.io/en/stable/reference/deprecation.html#deprecation-of-object-mode-fall-back-behaviour-when-using-jit for details.
@jit(parallel=True, nopython=False)
@jit(parallel=True, nopython=False)
读取矩阵数据
- 分析前,需要将 input.rds 转化为三个矩阵文件,才能读取
- 分析前,需要注释好细胞类型
python
adata = sc.read_10x_mtx("filtered_feature_bc_matrix/")
spatial = pd.read_csv('filtered_feature_bc_matrix/cell_locations.tsv',sep="\t",index_col=0)
spatial = spatial.loc[:,("x","y")]
selected_rows = spatial.loc[spatial.index.isin(adata.obs_names)]
selected_rows.columns = ["imagecol","imagerow"]
selected_rows = selected_rows.reindex(adata.obs_names)
# 使用st.create_stlearn可以将表达,空间位置导入anndata对象;若还想要导入图片,添加参数image_path。SeekSpace技术中一个像素点的大小是约为0.265385微米,故scale=0.265385。
a = st.create_stlearn(count=adata.to_df(),spatial=selected_rows,library_id="N1", scale=0.265385,spot_diameter_fullres=10)
cluster_name = "celltype"
celltype = pd.read_csv("../../data/AY1748480899609/meta.tsv",index_col=0,sep = "\t")
celltype = celltype.loc[a.obs.index]
## meta_celltype_colums_name为原rds里metadata中细胞类型列名
meta_celltype_colums_name = "CellAnnotation"
a.obs[cluster_name] = celltype[meta_celltype_colums_name]
a.obs[cluster_name] = a.obs[cluster_name].astype('category')
#a = a[a.obs["celltype"].isin(["HSC","Fibroblasts"])]
#a.layers["raw_count"] = a.X
a.layers["counts"] = a.X
sc.pp.normalize_total(a, target_sum=1e4)
sc.pp.log1p(a)
sc.pp.calculate_qc_metrics(a, percent_top=None, log1p=False, inplace=True)- Step 1: 使用半径搜索邻居
- Step 2: 处理邻居索引(按距离排序 + 排除自身)
- Step 3: 计算窗口和(处理稀疏区域)
- Step 4: 聚类邻域类型
python
def cell_neighbors(adata,column,radius=20,n_clusters=10):
spatial_coords = adata.obsm['spatial']
onehot_encoding = pd.get_dummies(adata.obs[column])
cluster_cols = adata.obs[column].unique()
values = onehot_encoding[cluster_cols].values
nbrs = NearestNeighbors(radius=radius,metric='euclidean').fit(spatial_coords)
distances,indices = nbrs.radius_neighbors(spatial_coords,return_distance=True)
sorted_indices = []
for i in range(len(indices)):
if len(indices[i]) == 0:
sorted_indices.append(np.array([]))
continue # 按距离排序并排除自身
sorted_order = np.argsort(distances[i])
neigh_indices = indices[i][sorted_order]
mask = (neigh_indices != i)
filtered_indices = neigh_indices[mask]
sorted_indices.append(filtered_indices)
def compute_window_sums(sorted_indices,values):
windows = []
for idx in range(len(sorted_indices)):
neighbors = sorted_indices[idx]
if len(neighbors) == 0: # 稀疏区域处理:使用自身类型填充
window_sum = values[idx] # 自身类型
else:
window_sum = values[neighbors].sum(axis=0)
# 归一化为比例分布
window_sum_norm = window_sum / (window_sum.sum() + 1e-6) # 防止零除
windows.append(window_sum_norm)
return np.array(windows)
windows = compute_window_sums(sorted_indices,values)
km = MiniBatchKMeans(n_clusters=n_clusters, random_state=0)
labels = km.fit_predict(windows)
adata.obs[f'CNs_{n_clusters}'] = [f'CN{i}' for i in labels] # 可视化 Fold Change
k_centroids = km.cluster_centers_
tissue_avgs = values.mean(axis=0)
fc = np.log2((k_centroids + 1e-6) / (tissue_avgs + 1e-6)) # 避免零除
fc_df = pd.DataFrame(fc,columns=cluster_cols)
fc_df.index = [f'CN{i}' for i in range(n_clusters)]
sns.set_style("white")
g = sns.clustermap(fc_df,vmin=-2,vmax=2,cmap="vlag",row_cluster=False,col_cluster=True,linewidths=0.5,figsize=(6,6))
g.ax_heatmap.tick_params(right=False,bottom=False)
return adata
cell_neighbors(adata=a,column="celltype")output
AnnData object with n_obs × n_vars = 20820 × 34506
obs: 'imagecol', 'imagerow', 'celltype', 'n_genes_by_counts', 'total_counts', 'CNs_10'
var: 'n_cells_by_counts', 'mean_counts', 'pct_dropout_by_counts', 'total_counts'
uns: 'spatial', 'log1p'
obsm: 'spatial'
layers: 'counts'
obs: 'imagecol', 'imagerow', 'celltype', 'n_genes_by_counts', 'total_counts', 'CNs_10'
var: 'n_cells_by_counts', 'mean_counts', 'pct_dropout_by_counts', 'total_counts'
uns: 'spatial', 'log1p'
obsm: 'spatial'
layers: 'counts'
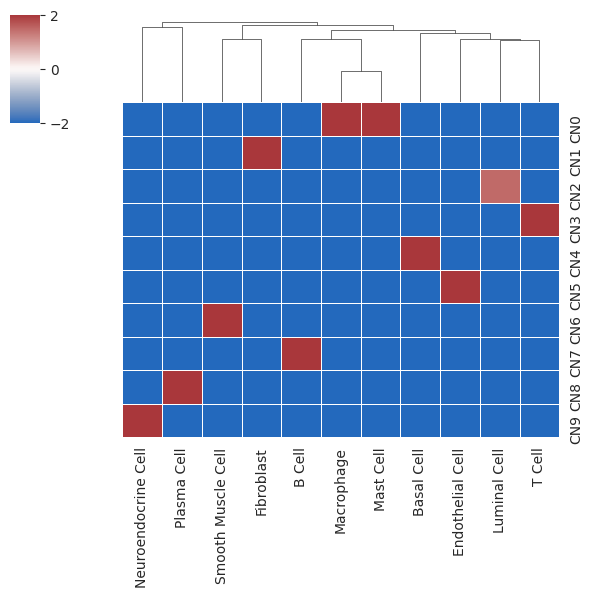
- 热图代表细胞类型在不同细胞 niche 的富集程度
- 横轴代表细胞类型;纵轴代表聚类的细胞 niche;中间的颜色代表富集程度python
aoutput
AnnData object with n_obs × n_vars = 20820 × 34506
obs: 'imagecol', 'imagerow', 'celltype', 'n_genes_by_counts', 'total_counts', 'CNs_10'
var: 'n_cells_by_counts', 'mean_counts', 'pct_dropout_by_counts', 'total_counts'
uns: 'spatial', 'log1p'
obsm: 'spatial'
layers: 'counts'
obs: 'imagecol', 'imagerow', 'celltype', 'n_genes_by_counts', 'total_counts', 'CNs_10'
var: 'n_cells_by_counts', 'mean_counts', 'pct_dropout_by_counts', 'total_counts'
uns: 'spatial', 'log1p'
obsm: 'spatial'
layers: 'counts'
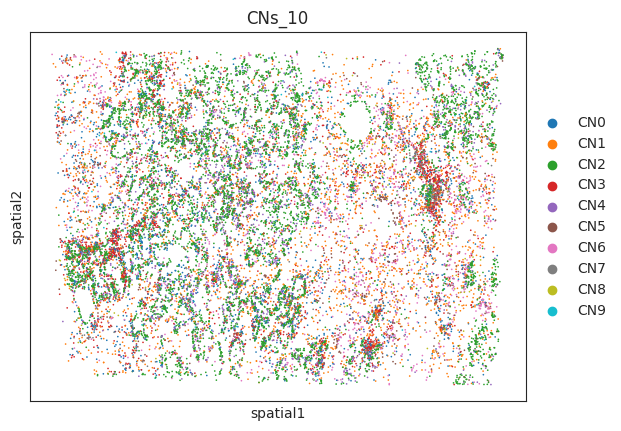
- 细胞 niche 在空间的可视化
python
sc.pl.embedding(a,"spatial",color=["CNs_10"])output
/PROJ2/FLOAT/jinwen/apps/miniconda3/envs/COMMOT/lib/python3.10/site-packages/scanpy/plotting/_tools/scatterplots.py:1235: FutureWarning: The default value of 'ignore' for the \`na_action\` parameter in pandas.Categorical.map is deprecated and will be changed to 'None' in a future version. Please set na_action to the desired value to avoid seeing this warning
color_vector = pd.Categorical(values.map(color_map))
color_vector = pd.Categorical(values.map(color_map))