SeekSpace 高级分析:空间细胞类型密度分布图绘制
时长: 8 分钟
字数: 1.6k 字
更新: 2026-02-28
阅读: 0 次
python
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.stats import gaussian_kde
from mpl_toolkits.mplot3d import Axes3D数据加载
先利用 R 从 rds 中保存空间坐标和注释结果
读取空间坐标
python
spatial_df = pd.read_csv("/PROJ2/FLOAT/weichendan/seekspace/SGS2401017/allsamples/00data/down_oss/ndGBM_5_2/ndGBM_5_2_barcode_centers.csv",index_col=0,sep=',')
spatial_df.drop(columns=['RNA_snn_res.0.8'], inplace=True)读取注释结果
python
#celltype_df = pd.read_csv('/PROJ2/FLOAT/weichendan/seekspace/SGS2401017/allsamples/08mistyR/ndGBM_5_2_celltype.csv', index_col=0)
celltype_df = pd.read_csv("/PROJ2/FLOAT/weichendan/seekspace/SGS2401017/updata_allsamples/00data/scanpy_obs.csv")
celltype_df = celltype_df.drop(celltype_df.columns[0], axis=1)
#celltype_df['new_cell_id'] = celltype_df['new_cell_id'].str.replace('_9', '')
celltype_df['new_cell_id'] = celltype_df['new_cell_id'].str.replace(r'_\d+$', '', regex=True)
celltype_df = celltype_df.set_index('new_cell_id')
celltype_df = celltype_df[celltype_df['sample_id'].str.contains("ndGBM_5_2")]python
spatial_df = spatial_df[spatial_df.index.isin(celltype_df.index)]
spatial_df = spatial_df.reindex(celltype_df.index)合并数据
python
merged_df = pd.merge(celltype_df, spatial_df, left_index=True, right_index=True,how="inner")python
merged_df| sample_id | cell_type | all_celltype | new_group | x | y | |
|---|---|---|---|---|---|---|
| new_cell_id | ||||||
| AGAGCCCCGATAGCTACCCTAATAACG | ndGBM_5_2 | Glioma | AC | GBM | 27913 | 15055 |
| CACCGTTATGGGAAACGTCCTATTAGG | ndGBM_5_2 | Fibroblast | Fibroblast | GBM | 29920 | 4106 |
| GCCATTCTACGCCTTTGCTAAGTACGA | ndGBM_5_2 | Glioma | AC | GBM | 34538 | 1969 |
| TAAGTCGCGACATCCGGCTAAGTACGA | ndGBM_5_2 | Oligodendrocyte | Oligodendrocyte | GBM | 27680 | 10535 |
| TACGTCCTACGAAGTGGGGACTGTGGA | ndGBM_5_2 | Fibroblast | Fibroblast | GBM | 43135 | 11168 |
| ... | ... | ... | ... | ... | ... | ... |
| CGCGTGACGATGCAAGGAAGGCACTAT | ndGBM_5_2 | Endothelial_cell | Endothelial_cell | GBM | 33528 | 12544 |
| GCCAGGTATGAGTCCGGGGACTGTGGA | ndGBM_5_2 | Endothelial_cell | Endothelial_cell | GBM | 40550 | 7503 |
| TGGGTTAATGACGTCGAAAGGCACTAT | ndGBM_5_2 | Endothelial_cell | Endothelial_cell | GBM | 20952 | 13660 |
| CCCGGAACGAGTCAGCCCCTAATAACG | ndGBM_5_2 | Endothelial_cell | Endothelial_cell | GBM | 41631 | 16713 |
| TCAGGGCGCTTTCTATCAGCGACGGAT | ndGBM_5_2 | Endothelial_cell | Endothelial_cell | GBM | 34379 | 11688 |
14055 rows × 6 columns
绘制 3D 空间密度图
以细胞类型 AC 为例,绘制其 3D 空间密度图
python
merged_df1 = merged_df[merged_df['all_celltype']=="AC"][['x', 'y']]
# 计算密度分布
xy = np.vstack([merged_df1.x, merged_df1.y])
kde = gaussian_kde(xy, bw_method=0.2)
xmin, xmax = merged_df1.x.min(), merged_df1.x.max()
ymin, ymax = merged_df1.y.min(), merged_df1.y.max()
xgrid = np.linspace(xmin, xmax, 200)
ygrid = np.linspace(ymin, ymax, 200)
X, Y = np.meshgrid(xgrid, ygrid)
Z = kde(np.vstack([X.ravel(), Y.ravel()])).reshape(X.shape)
# 创建画布
fig = plt.figure(figsize=(15, 5))
# ---------------------------
# 3D密度图(关键视角设置)
# ---------------------------
ax3d = fig.add_subplot(111, projection='3d')
###设置长:宽:高
ax3d.set_box_aspect([15, 5, 5])
## 取消网格
ax3d.grid(False)
# 绘制密度曲面
surf = ax3d.plot_surface(X, Y, Z, cmap='RdYlGn_r',
rstride=5, cstride=5,
alpha=1, antialiased=True,
edgecolor='none')
# 视角参数设置(关键调整)
ax3d.view_init(elev=25, azim=-90) # 通过azim调整坐标轴投影方向
#ax3d.set_proj_type('ortho') # 正交投影消除透视变形
# 坐标轴范围同步
ax3d.set_xlim(xmin, xmax)
ax3d.set_ylim(ymin, ymax)
ax3d.set_zlim(0, Z.max()*1.1)
# 取消坐标轴刻度
ax3d.set_xticks([])
ax3d.set_yticks([])
ax3d.set_zticks([])
# 取消坐标轴线
ax3d.xaxis.line.set_color((0,0,0,0))
ax3d.yaxis.line.set_color((0,0,0,0))
ax3d.zaxis.line.set_color((0,0,0,0))
# 取消坐标轴标签
ax3d.set_xlabel('')
ax3d.set_ylabel('')
ax3d.set_zlabel('')
# 隐藏冗余标签
ax3d.set_xticklabels([])
ax3d.set_yticklabels([])
#ax3d.set_zlabel('Density', labelpad=15, fontsize=12)
# 背景和三面墙变成白色
ax3d.set_facecolor('white')
ax3d.xaxis.set_pane_color((1.0, 1.0, 1.0, 1.0))
ax3d.yaxis.set_pane_color((1.0, 1.0, 1.0, 1.0))
ax3d.zaxis.set_pane_color((1.0, 1.0, 1.0, 1.0))
plt.subplots_adjust(left=0, right=1, top=1, bottom=0)
#plt.savefig('output.png', dpi=300)
plt.show()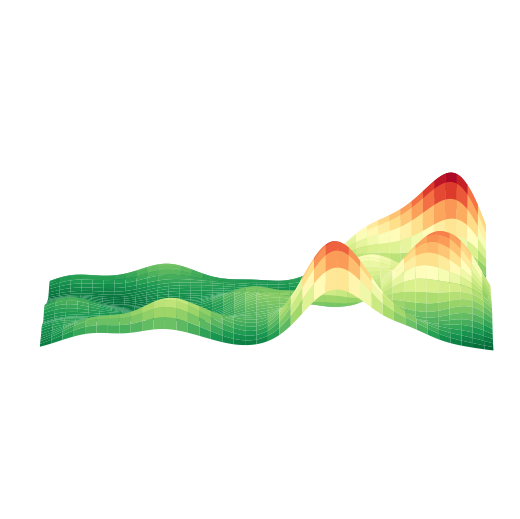
绘制 3D 空间密度图的映射图
以细胞类型 AC 为例,绘制其 3D 空间密度图的映射图
python
merged_df1 = merged_df[merged_df['all_celltype']=="AC"][['x', 'y']]
# 计算密度分布
xy = np.vstack([merged_df1.x, merged_df1.y])
kde = gaussian_kde(xy, bw_method=0.2)
xmin, xmax = merged_df1.x.min(), merged_df1.x.max()
ymin, ymax = merged_df1.y.min(), merged_df1.y.max()
xgrid = np.linspace(xmin, xmax, 200)
ygrid = np.linspace(ymin, ymax, 200)
X, Y = np.meshgrid(xgrid, ygrid)
Z = kde(np.vstack([X.ravel(), Y.ravel()])).reshape(X.shape)
# 创建画布
fig = plt.figure(figsize=(15, 5))
# ---------------------------
# 3D密度图(关键视角设置)
# ---------------------------
ax3d = fig.add_subplot(111, projection='3d')
###设置长:宽:高
ax3d.set_box_aspect([15, 5, 0.1])
## 取消网格
ax3d.grid(False)
# 绘制密度曲面
surf = ax3d.plot_surface(X, Y, Z, cmap='RdYlGn_r',
rstride=5, cstride=5,
alpha=1, antialiased=True,
edgecolor='none')
# 视角参数设置(关键调整)
ax3d.view_init(elev=25, azim=-90) # 通过azim调整坐标轴投影方向
#ax3d.set_proj_type('ortho') # 正交投影消除透视变形
# 坐标轴范围同步
ax3d.set_xlim(xmin, xmax)
ax3d.set_ylim(ymin, ymax)
ax3d.set_zlim(0, Z.max()*1.1)
# 取消坐标轴刻度
ax3d.set_xticks([])
ax3d.set_yticks([])
ax3d.set_zticks([])
# 取消坐标轴线
ax3d.xaxis.line.set_color((0,0,0,0))
ax3d.yaxis.line.set_color((0,0,0,0))
ax3d.zaxis.line.set_color((0,0,0,0))
# 取消坐标轴标签
ax3d.set_xlabel('')
ax3d.set_ylabel('')
ax3d.set_zlabel('')
# 隐藏冗余标签
ax3d.set_xticklabels([])
ax3d.set_yticklabels([])
#ax3d.set_zlabel('Density', labelpad=15, fontsize=12)
# 背景和三面墙变成白色
ax3d.set_facecolor('white')
ax3d.xaxis.set_pane_color((1.0, 1.0, 1.0, 1.0))
ax3d.yaxis.set_pane_color((1.0, 1.0, 1.0, 1.0))
ax3d.zaxis.set_pane_color((1.0, 1.0, 1.0, 1.0))
plt.subplots_adjust(left=0, right=1, top=1, bottom=0)
#plt.savefig('output.png', dpi=300)
plt.show()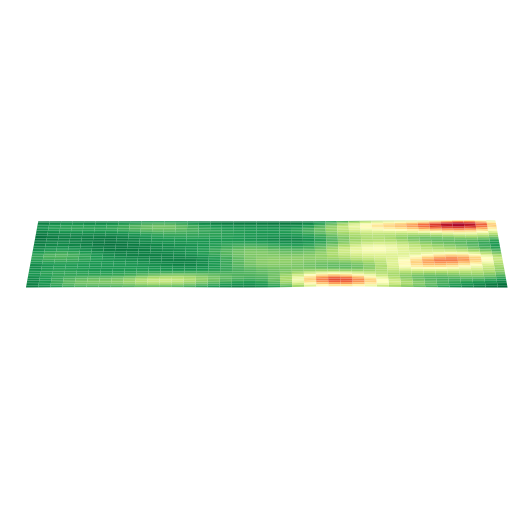
绘制 2D 密度图
python
fig2d, ax2d = plt.subplots(figsize=(6, 4))
contour = ax2d.contourf(X, Y, Z, levels=20, cmap='RdYlGn_r')
# 坐标轴设置
ax2d.set(xlim=(xmin, xmax), ylim=(ymin, ymax), aspect='equal')
# 取消坐标轴刻度
ax2d.set_xticks([])
ax2d.set_yticks([])
# 取消坐标轴标签
ax2d.set_xlabel('')
ax2d.set_ylabel('')
plt.show()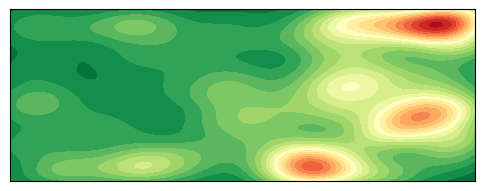
python
python
